Latent variable modelling for massive and complex datasets
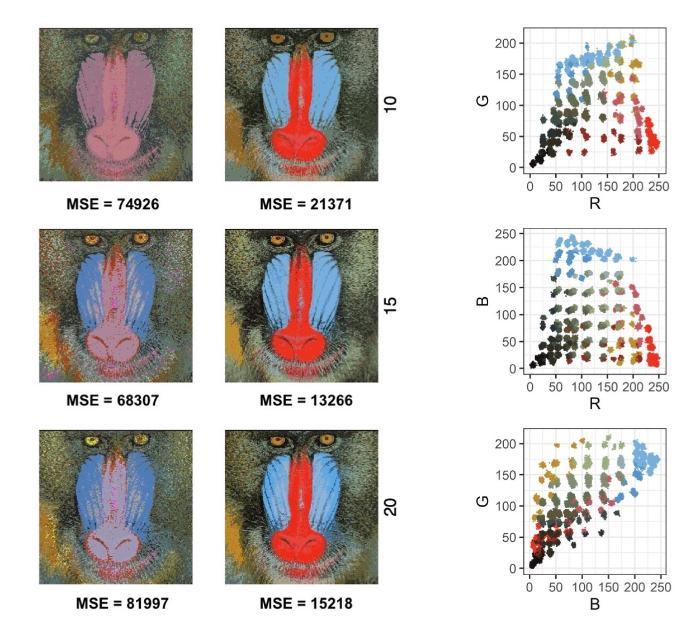
Massive advances in genomic sequencing-based and imaging technologies in the last two decades have generated the potential to make novel biological discoveries at extremely high resolutions- but have also led to numerous challenging problems in how to sensibly and accurately analyse the generated datasets. These data are typically of large dimensions- leading to computational obstacles; are subject to various technical artefacts; and their distributions exhibit complex features, such as long-ranging correlations, non-ellipsoidal shapes, skewness and multimodality, which cause difficulties in making successful inference through classical standard statistical models. We have been developing latent variable-based Bayesian hierarchical modelling approaches for clustering in complex datasets, that lead to efficient and powerful computational methods in enabling inference and biological discovery. One example involves clustering high-volume genotyping data- finding subgroups with common features is often a necessary first step with the downstream goal of detection of genetic variants associated with specific health outcomes.

Researchers
Publications
- Bayesian hierarchical mixture models for detecting non‐normal clusters applied to noisy genomic and environmental datasets, Australian & New Zealand Journal of Statistics, 64(2), 313-337 (2022).
- Bayesian modeling of factorial time-course data with applications to a bone aging gene expression study, Journal of Applied Statistics, 48(10), 1730-1754 (2021).
- Bayesian clustering of skewed and multimodal data using geometric skewed normal distributions, Computational Statistics & Data Analysis, 152, 107040 (2020).
Organisations

