The Jarrett Group
Our group has a longstanding interest in the role of viruses in leukaemia and lymphoma. Much of our research is focused on Hodgkin lymphoma, one of the commonest malignancies affecting young adults.
Research programme
The Epstein-Barr virus (EBV) is associated with a proportion of cases of Hodgkin lymphoma, and we believe that this association is causal. In these ‘EBV-positive’ cases, EBV is present in all the tumour cells, the viral infection is clonal, and viral gene products with plausible biological function are expressed.
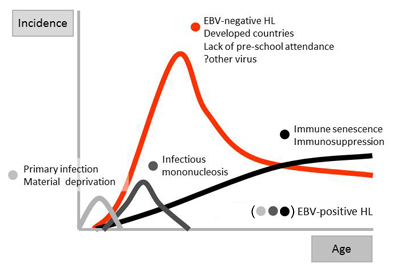 EBV-positive and negative cases show distinct epidemiological features and we have put forward the four disease model of Hodgkin lymphoma, which divides the disease into subgroups on the basis of EBV-positivity, age at diagnosis, and age at infection with EBV. The aetiology of EBV-negative Hodgkin lymphoma is obscure although we believe that another, hitherto unidentified, virus may be involved. The search for this virus has been, and continues to be, a major interest.
EBV-positive and negative cases show distinct epidemiological features and we have put forward the four disease model of Hodgkin lymphoma, which divides the disease into subgroups on the basis of EBV-positivity, age at diagnosis, and age at infection with EBV. The aetiology of EBV-negative Hodgkin lymphoma is obscure although we believe that another, hitherto unidentified, virus may be involved. The search for this virus has been, and continues to be, a major interest.
As part of international collaborations, we have shown that genetic risk factors for EBV-positive and negative disease are largely distinct. The strongest associations for both disease subtypes are with HLA alleles and SNPs in the MHC region; our most recent analysis (Johnson et al, 2015) provides further evidence that HLA-A*01:01 is associated with an increased risk of EBV-associated Hodgkin lymphoma, and that the SNP rs6903608 is the strongest predictor of risk of EBV-negative disease. Prior infectious mononucleosis is also associated with an increased risk of EBV-positive Hodgkin lymphoma, and this appears independent of HLA associations.
Current research
Hodgkin lymphoma
 Our research on EBV-positive Hodgkin lymphoma is focused on investigating the biology underlying associations with specific HLA alleles and with infectious mononucleosis. For EBV-negative Hodgkin lymphoma we are using next generation sequencing approaches to look for a novel virus in tumour cells enriched from biopsy samples. In translational studies, we are investigating biomarkers, including CCL17, which appear promising in the early diagnosis of Hodgkin lymphoma and in assessing response to treatment.
Our research on EBV-positive Hodgkin lymphoma is focused on investigating the biology underlying associations with specific HLA alleles and with infectious mononucleosis. For EBV-negative Hodgkin lymphoma we are using next generation sequencing approaches to look for a novel virus in tumour cells enriched from biopsy samples. In translational studies, we are investigating biomarkers, including CCL17, which appear promising in the early diagnosis of Hodgkin lymphoma and in assessing response to treatment.
Canine lymphoma
We are also applying next generation sequencing approaches to the study of canine lymphoma, which shows remarkable similarities to non-Hodgkin lymphoma in man. The data are being used to investigate viral involvement, to identify gene expression signatures in lymphoma subtypes, and to identify driver mutations in tumour cells.
Human herpesvirus 6
Human herpesvirus 6 (HHV6) A and B are unusual among herpesviruses in that they integrate into telomere sequences at the ends of chromosomes. In 1-3% of the population, HHV6A or B is integrated in every cell in the body and passed to subsequent generations in a Mendelian fashion; however, little is known about the consequences of this so-called inherited chromosomally integrated HHV6 (iciHHV6). We are investigating the prevalence and disease associations of iciHHV6 in a large, population-based, family study - the Generation Scotland study. In collaborative studies, we are also investigating the sequence and evolution of HHV6 genomes.
Indentification of iciHHV-6 using droplet digital PCR.
Research group members
|
Natasha Jesudason |
Annette Lake |
Michael Wood |
Collaborators
Internal
- Dr Andrew Davison – HHV6 genomics
- Dr Robert Gifford – HHV6 evolution
- Dr Jo Morris – canine lymphoma
External
- Dr Henrik Hjalgrim (Statens Serum Institute, Copenhagen) – Hodgkin lymphoma, EBV
- Prof Dr Anke van den Berg (University Medical Center Groningen) – Hodgkin lymphoma, EBV
- Dr Arjan Diepstra (University Medical Center Groningen) - Hodgkin lymphoma, EBV
- Dr James McKay (International Agency for Research on Cancer, Lyon) - Hodgkin lymphoma, GWAS
- Dr Wendy Cozen (Keck School of Medicine, University of Southern California) - Hodgkin lymphoma, EBV, GWAS
- Prof Richard Houlston (Institute of Cancer Research, Sutton) - Hodgkin lymphoma, GWAS
- Dr Jonathan Fromm (Seattle Cancer Care Alliance) – enrichment of Hodgkin lymphoma tumour cells
- Dr Nicola Royle (University of Leicester) – HHV6




