Molecular Pharmacology & Cellular Signalling
Our research looks at the molecular mechanisms that underpin the basis of physiology and the development of disease.
Staff
Publications
2026
Xia, L., Alvim, J. C., Nguyen, T.-H., Lefoulon, C., Silva-Alvim, F. A. L., Yu, Z., Klejchova, M., Farami, S., Waghmare, S., Karnik, R., Blatt, M. R. (2026) A guard cell carbonic anhydrase binds and regulates SLAC1 separate from its catalytic activity. Nature Communications, 17, (doi: 10.1038/s41467-026-70596-9)
Yu, Z., Waghmare, S., Farami, S., Blatt, M. R., Karnik, R. (2026) CO2 - sensitive K+ channel traffic affects stomata and whole plant water-use. Journal of Integrative Plant Biology, (doi: 10.1111/jipb.70269)
Malone, M. A., Yin, R., Li, Y., Jenkins, L., Guseinov, A.-A., Marsango, S., Huggett, M., Huggett, M., Boyle, A., Morrison, A., Tikhonova, I. G., Milligan, G., Jamieson, A. G. (2026) Discovery, structure–activity relationship, and functional characterization of a chromenopyrrole series as orthosteric antagonists of GPR84. Journal of Medicinal Chemistry, 69, pp. 7945-7965. (doi: 10.1021/acs.jmedchem.5c03367)
Fleming, B., Guseinov, A.-A., Tikhonova, I. G., Perry-Hauser, N., Milligan, G. (2026) Steroid recognition in adhesion GPCRs: a structural and pharmacological perspective. Pharmacology Research and Perspectives, 14, (doi: 10.1002/prp2.70221)
Brettell, S. B., Fuentes-Guerra Bustos, C., Sharma, S., Cann, G., Carruthers, L. V., Begen, A., Milligan, G., Clarke, D. J., Tobin, A. B., Jamieson, A. (2026) Shaping antimalarials: a geometry-first approach to PfCLK3 covalent inhibitors. Journal of Medicinal Chemistry, 69, pp. 3378-3395. (doi: 10.1021/acs.jmedchem.5c03342)
Blatt, M. R., Hills, A., Lawson, T., Magill, J. (2026) Cell wall water shields stomata against falling leaf airspace humidity. New Phytologist, (doi: 10.1111/nph.70998)
Rudzik, N., Stubbs, F., Watterson, K., McVey, M. (2026) Skills auditing, surfacing and signalling: embedding employability focussed activities in a Level 1 Life Sciences course.
Silva-Alvim, F. A.L., Piro, L., Santelia, D., Blatt, M. R. (2026) Guard cell starch and malate metabolism facilitate stomatal opening in response to low CO₂ New Phytologist, 249, pp. 189-197. (doi: 10.1111/nph.70639)
2025
Alshuweishi, Y., Alghamdi, F. B., Patrick, K., Salt, I. P. (2025) AICAR inhibits insulin-stimulated glucose uptake in 3T3-L1 adipocytes via an AMPK-independent, ZMP-dependent mechanism. Cells, 14, (doi: 10.3390/cells14221811)
Berrocal-Martin, R., Sansom, H. G., Orosz, K., Paterson, M. B., Smith, B. O., Magennis, S. W. (2025) Time-resolved fluorescence detection of nicked DNA via site-specific stacking of sulfo-Cy3: the role of charge, polarity, linker and sequence context. Journal of Physical Chemistry B, 129, pp. 11626-11635. (doi: 10.1021/acs.jpcb.5c03845)
Nakasone, M., Buetow, L., Gabrielsen, M., Ahmed, S. F., Majorek, K. A., Sibbet, G. J., Smith, B. O., Huang, D. T. (2025) Tuning ubiquitin transfer by RING E3 ubiquitin ligases through the linchpin residue. Life Science Alliance, 8, (doi: 10.26508/lsa.202503394)
Brettell, S. B., Cann, G., Begen, A., Sharma, S., Mahindra, A., Carruthers, L. V., Milligan, G., Clarke, D. J., Tobin, A. B., Jamieson, A. G. (2025) Lysine targeting covalent inhibitors of malarial kinase PfCLK3. RSC Medicinal Chemistry, 16, pp. 3530-3540. (doi: 10.1039/d5md00335k)
Zhang, C., Zhang, X., Guseinov, A.-A., Jenkins, L., Valentini, A., Marsango, S., Schultz-Knudsen, K., Ulven, T., Rexen Ulven, E., Tikhonova, I. G., Milligan, G., Zhang, C. (2025) Allosteric modulation and biased signalling at free fatty acid receptor 2. Nature, 643, pp. 1428-1438. (doi: 10.1038/s41586-025-09186-6)
Valentini, A., Dibnah, B., Ciba, M., Duncan, E. M., Manandhar, A., Strellis, B., Vita, L., Lucianno, O., Massey, C., Coe, S., Ulven, T., Hudson, B. D., Ulven, E. R. (2025) Multicolored sequential resonance energy transfer for detection of simultaneous ligand binding at G protein-coupled receptors. Nature Communications, 16, (doi: 10.1038/s41467-025-61690-5)
O’Brien, S. L., Tripp, E., Barki, N., Blondel-Tepaz, E., Smith, G., Boufersaoui, A., Roberts, J., Pike, J. A., Correia, J., Miljus, T., Bouvier, M., Tennant, D. A., Hudson, B. D., Gerhart-Hines, Z., Milligan, G., Schwartz, T. W., Calebiro, D. (2025) Intracrine FFA4 signaling controls lipolysis at lipid droplets. Nature Chemical Biology, (doi: 10.1038/s41589-025-01982-5)
Li, H., Xu, L., Venkata, S. P. M., Minjares, M., Melhem, H., Kowluru, A., Niess, J. H., Milligan, G., Wang, J.-M. (2025) G protein-coupled receptor 35 suppresses oxidative stress response 1 in diabetic wound healing. Diabetes, 74, pp. 1233-1246. (doi: 10.2337/db24-0737)
Alshammari, W.S., Duncan, E.M., Vita, L., Kenawy, M., Dibnah, B., Wabitsch, M., Gould, G.W., Hudson, B.D. (2025) Inverse agonism of the FFA4 free fatty acid receptor controls both adipogenesis and mature adipocyte function. Cellular Signalling, 131, (doi: 10.1016/j.cellsig.2025.111714)
Ralston, M. R., McCreath, G., Lees, Z. J., Salt, I. P., Sim, M. A.B., Watson, M. J., Freeman, D. J. (2025) Beyond body mass index: exploring the role of visceral adipose tissue in intensive care unit outcomes. BJA Open, 14, (doi: 10.1016/j.bjao.2025.100391)
Gonzalez-Valdivieso, J., Ciccone, G., Dhawan, U., Quon, T., Barcelona-Estaje, E., Rodrigo-Navarro, A., Castillo, R. R., Milligan, G., Rico, P., Salmeron-Sanchez, M. (2025) NaBC1 boron transporter enables myoblast response to substrate rigidity via fibronectin-binding integrins. Advanced Science, 12, (doi: 10.1002/advs.202407548)
Zhang, X., Carroll, W., Binh-Anh, T., Nguyen, T.-H., Yang, Z., Ma, M., Huang, X., Hills, A., Guo, H., Karnik, R., Blatt, M. R., Zhang, P. (2025) GORK K+ channel structure and gating vital to informing stomatal engineering. Nature Communications, 16, (doi: 10.1038/s41467-025-57287-7)
McAleese, C., Joudah, G., Salt, I.P., Petrie, J.R., Leiper, J.M., Dowsett, L. B. (2025) Heterogeneous metabolic response of endothelial cells from different vascular beds to experimental hyperglycaemia and metformin. Journal of Physiology, (doi: 10.1113/JP288006)
Alvim, F. L. S., Alvim, J., Hibberd, J. M., Harvey, A. R., Blatt, M. R. (2025) A C4 plant K+ channel accelerates stomata to enhance C3 photosynthesis and water use efficiency. Plant Physiology, 197, (doi: 10.1093/plphys/kiaf039)
Nguyen, T.-H., Krasauskas, J., Nguyen, T. B.-A., Noureen, A., Smedley, M., Christie, J. M., Harwood, W., Blatt, M. R., Hundleby, P. (2025) A promoter collection for cell-targeted analysis within the stomatal complex. Plant Direct, 9, (doi: 10.1002/pld3.70045)
Quon, T., Lin, L.-C., Ganguly, A., Hudson, B. D., Tobin, A. B., Milligan, G. (2025) Biased constitutive signaling of the G protein-coupled receptor GPR35 suppresses gut barrier permeability. Journal of Biological Chemistry, 301, (doi: 10.1016/j.jbc.2024.108035)
Waghmare, S., Xia, L., Phan Ly, T., Xu, J., Farami, S., Burchmore, R., Blatt, M. R., Karnik, R. (2025) SYNTAXIN OF PLANTS 132 underpins secretion of cargoes associated with salicylic acid signaling and pathogen defense. Plant Physiology, 197, (doi: 10.1093/plphys/kiae541)
2024
Adler, L., Sing Lau, C., Shaikh, K. M., van Maldegem, K. A., Payne-Dwyer, A. L., Lefoulon, C., Girr, P., Atkinson, N., Barrett, J., Emrich-Mills, T. Z., Dukic, E., Blatt, M. R., Leake, M. C., Peltier, G., Spetea, C., Burlacot, A., McCormick, A. J., Mackinder, L. C.M., Walker, C. E. (2024) Bestrophin-like protein 4 is involved in photosynthetic acclimation to light fluctuations in Chlamydomonas. Plant Physiology, 196, pp. 2374-2394. (doi: 10.1093/plphys/kiae450)
Lefoulon, C., Blatt, M. R. (2024) Guard cell K+ channels of Kalanchoë follow the diel cycle of crassulacean acid metabolism. Plant Physiology, 196, pp. 2300-2303. (doi: 10.1093/plphys/kiae506)
Hwej, A., Al-Ferjani, A., Alshuweishi, Y., Naji, A., Kennedy, S., Salt, I. (2024) Lack of AMP-activated protein kinase-α1 reduces nitric oxide synthesis in thoracic aorta perivascular adipose tissue. Vascular Pharmacology, 157, (doi: 10.1016/j.vph.2024.107437)
Brettell, S. B., Janha, O., Begen, A., Cann, G., Sharma, S., Olaniyan, N., Yelland, T., Hole, A. J., Alam, B., Mayville, E., Gillespie, R., Capper, M., Fidock, D. A., Milligan, G., Clarke, D. J., Tobin, A. B., Jamieson, A. G. (2024) Targeting PfCLK3 with covalent inhibitors: a novel strategy for malaria treatment. Journal of Medicinal Chemistry, 67, pp. 18895-18910. (doi: 10.1021/acs.jmedchem.4c01300)
Salt, I. P., Carling, D. (2024) A special issue of Essays in Biochemistry on AMPK and AMPK-related kinases. Essays In Biochemistry, 68, pp. 269-271. (doi: 10.1042/EBC20240038)
Dearlove, E. L., Chatrin, C., Buetow, L., Ahmed, S. F., Schmidt, T., Bushell, M., Smith, B. O., Huang, D. T. (2024) DTX3L ubiquitin ligase ubiquitinates single-stranded nucleic acids. eLife, 13, (doi: 10.7554/elife.98070)
Nguyen, T.-H., Blatt, M. R. (2024) Surrounded by luxury: the necessities of subsidiary cells. Plant, Cell and Environment, 47, pp. 3316-3329. (doi: 10.1111/pce.14872)
Su, J., He, B., Li, P., Yu, B., Cen, Q., Xia, L., Jing, Y., Wu, F., Karnik, R., Xue, D., Blatt, M. R., Wang, Y. (2024) Overexpression of tonoplast Ca2+‐ATPase in guard cells synergistically enhances stomatal opening and drought tolerance. Journal of Integrative Plant Biology, 66, pp. 1587-1602. (doi: 10.1111/jipb.13721)
Blatt, M. R. (2024) A charged existence: a century of transmembrane ion transport in plants. Plant Physiology, 195, pp. 79-110. (doi: 10.1093/plphys/kiad630)
Baena, G., Xia, L., Waghmare, S., Yu, Z., Guo, Y., Blatt, M. R., Zhang, B., Karnik, R. (2024) Arabidopsis SNARE SYP132 impacts on PIP2;1 trafficking and function in salinity stress. Plant Journal, 118, pp. 1036-1053. (doi: 10.1111/tpj.16649)
Milligan, G. (2024) Editorial for GPR84 Pharmacology. British Journal of Pharmacology, 181, pp. 1497-1499. (doi: 10.1111/bph.16340)
Marsango, S., Milligan, G. (2024) Regulation of the pro-inflammatory G protein-coupled receptor GPR84. British Journal of Pharmacology, 181, pp. 1500-1508. (doi: 10.1111/bph.16098)
Pal, S., Nare, Z., Rao, V. A., Smith, B. O., Morrison, I., Fitzgerald, E. A., Scott, A., Bingham, M. J., Pesnot, T. (2024) Accelerating BRPF1b hit identification with BioPhysical and Active Learning Screening (BioPALS) ChemMedChem, 19, (doi: 10.1002/cmdc.202300590)
Silva-Alvim, F. A.L., Alvim, J. C., Harvey, A., Blatt, M. R. (2024) Speedy stomata of a C4 plant correlate with enhanced K+ channel gating. Plant, Cell and Environment, 47, pp. 817-831. (doi: 10.1111/pce.14775)
Tikhonova, I., Zhang, X., Guseinov, A.-A., Jenkins, L., Li, K., Milligan, G., Zhang, C. (2024) Structural basis for the ligand recognition and signaling of free fatty acid receptors. Science Advances, 10, (doi: 10.1126/sciadv.adj2384)
2023
Barki, N., Jenkins, L., Marsango, S., Dedeo, D., Bolognini, D., Dwomoh, L., Abdelmalik, A. M., Nilsen, M., Stoffels, M., Nagel, F., Schulz, S., Tobin, A. B., Milligan, G. (2023) Phosphorylation bar-coding of free fatty acid receptor 2 is generated in a tissue-specific manner. eLife, 12, (doi: 10.7554/eLife.91861)
Kalita, M., Park, J. H., Kuo, R. C., Hayee, S., Marsango, S., Straniero, V., Alam, I. S., Rivera-Rodriguez, A., Pandrala, M., Carlson, M. L., Reyes, S. T., Jackson, I. M., Suigo, L., Luo, A., Nagy, S. C., Valoti, E., Milligan, G., Habte, F., Shen, B., James, M. L. (2023) PET imaging of innate immune activation using 11C radiotracers targeting GPR84. JACS Au, 3, pp. 3297-3310. (doi: 10.1021/jacsau.3c00435)
Jaffery, H., Huesa, C., Chilaka, S., Cole, J., Doonan, J., Akbar, M., Dunning, L., Tanner, K. E., van 't Hof, R. J., McInnes, I. B., Carmody, R. J., Goodyear, C. S. (2023) IĸB Protein BCL3 as a controller of osteogenesis and bone health. Arthritis and Rheumatology, 75, pp. 2148-2160-2148-2160. (doi: 10.1002/art.42639)
Nguyen, T. B.-A., Lefoulon, C., Nguyen, T.-H., Blatt, M. R., Carroll, W. (2023) Engineering stomata for enhanced carbon capture and water-use efficiency. Trends in Plant Science, 28, pp. 1290-1309. (doi: 10.1016/j.tplants.2023.06.002)
Nguyen, T.-H., Silva-Alvim, F. A. L., Hills, A., Blatt, M. R. (2023) OnGuard3e: a predictive, ecophysiology‐ready tool for gas exchange and photosynthesis research. Plant, Cell and Environment, 46, pp. 3644-3658. (doi: 10.1111/pce.14674)
Irshad, Z., Lund, J., Sillars, A., Løvsletten, N. G., Gharanei, S., Salt, I. P., Freeman, D. J., Gill, J. M.R., Thoresen, G. H., Rustan, A. C., Zammit, V. A. (2023) The roles of DGAT1 and DGAT2 in human myotubes are dependent on donor patho-physiological background. FASEB Journal, 37, (doi: 10.1096/fj.202300960RR)
Ganguly, A., Quon, T., Jenkins, L., Joseph, B., Al-awar, R., Chevigne, A., Tobin, A. B., Uehling, D. E., Hoffmann, C., Drube, J., Milligan, G. (2023) G protein-receptor kinases 5/6 are the key regulators of G protein-coupled receptor 35-arrestin interactions. Journal of Biological Chemistry, 299, (doi: 10.1016/j.jbc.2023.105218)
Zhang, X., Wang, Y., Supekar, S., Cao, X., Zhou, J., Dang, J., Chen, S., Jenkins, L., Marsango, S., Li, X., Liu, G., Milligan, G., Feng, M., Fan, H., Gong, W., Zhang, C. (2023) Pro-phagocytic function and structural basis of GPR84 signaling. Nature Communications, 14, (doi: 10.1038/s41467-023-41201-0)
Ciba, M., Dibnah, B., Hudson, B. D., Ulven, E. R. (2023) Development and characterization of potent succinate receptor fluorescent tracers. Journal of Medicinal Chemistry, 66, pp. 8951-8974. (doi: 10.1021/acs.jmedchem.3c00552)
Duncan, E., Vita, L., Dibnah, B., Hudson, B. D. (2023) Metabolite sensing GPCRs controlling interactions between adipose tissue and inflammation. Frontiers in Endocrinology, 14, (doi: 10.3389/fendo.2023.1197102)
Karnik, R., Blatt, M. R. (2023) Analyzing protein–protein interactions using the split-ubiquitin system. Humana Press
Dowsett, L., Duluc, L., Higgins, E., Alghamdi, F., Fast, W., Salt, I. P., Leiper, J. (2023) Asymmetric dimethylarginine positively modulates calcium-sensing receptor signalling to promote lipid accumulation. Cellular Signalling, 107, (doi: 10.1016/j.cellsig.2023.110676)
Watterson, K., Daly, C., Uwase, C., MacRobbie, C. (2023) The Development of a Novel Educational Resource to Support Student Learning and Engagement in Pharmacology. (doi: 10.1111/bph.16108)
Deni, I. et al. (2023) Mitigating the risk of antimalarial resistance via covalent dual-subunit inhibition of the Plasmodium proteasome. Cell Chemical Biology, 30, pp. 470-485.e6. (doi: 10.1016/j.chembiol.2023.03.002)
Valentini, A., Schultz-Knudsen, K., Højgaard Hansen, A., Tsakoumagkou, A., Jenkins, L., Christensen, H. B., Manandhar, A., Milligan, G., Ulven, T., Rexen Ulven, E. (2023) Discovery of potent tetrazole free fatty acid receptor 2 antagonists. Journal of Medicinal Chemistry, 66, pp. 6105-6121. (doi: 10.1021/acs.jmedchem.2c01935)
Milligan, G. (2023) GPR35: from enigma to therapeutic target. Trends in Pharmacological Sciences, 44, pp. 263-273. (doi: 10.1016/j.tips.2023.03.001)
Javitt, A. et al. (2023) The proteasome regulator PSME4 modulates proteasome activity and antigen diversity to abrogate antitumor immunity in NSCLC. Nature Cancer, 4, pp. 629-647. (doi: 10.1038/s43018-023-00557-4)
Adam, S., Zheng, D., Klein, A., Volz, C., Mullen, W., Shirran, S. L., Smith, B. O., Kalinina, O. V., Müller, R., Koehnke, J. (2023) Unusual peptide-binding proteins guide pyrroloindoline alkaloid formation in crocagin biosynthesis. Nature Chemistry, 15, pp. 560-568. (doi: 10.1038/s41557-023-01153-w)
Meda Venkata, S. P., Li, H., Xu, L., Koh, J. Y., Nguyen, H., Minjares, M., Li, C., Kowluru, A., Milligan, G., Wang, J.-M. (2023) Inhibition of GPR39 restores defects in endothelial cell–mediated neovascularization under the duress of chronic hyperglycemia: evidence for regulatory roles of the sonic hedgehog signaling axis. Proceedings of the National Academy of Sciences of the United States of America, 120, pp. e2208541120. (doi: 10.1073/pnas.2208541120)
2022
Seale, M., Zhdanov, O., Soons, M. B., Cummins, C., Kroll, E., Blatt, M. R., Zare-Behtash, H., Busse, A., Mastropaolo, E., Bullock, J. M., Viola, I. M., Nakayama, N. (2022) Environmental morphing enables informed dispersal of the dandelion diaspore. eLife, 11, (doi: 10.7554/eLife.81962)
Kowalczyk, D., Nakasone, M. A., Smith, B. O., Huang, D. T. (2022) Bivalent binding of p14ARF to MDM2 RING and acidic domains inhibits E3 ligase function. Life Science Alliance, 5, (doi: 10.26508/lsa.202201472)
Bremner, S. K., Al Shammari, W. O., Milligan, R. S., Hudson, B. D., Sutherland, C., Bryant, N. J., Gould, G. W. (2022) Pleiotropic effects of Syntaxin16 identified by gene editing in cultured adipocytes. Frontiers in Cell and Developmental Biology, 10, (doi: 10.3389/fcell.2022.1033501)
Horaruang, W., Klejchová, M., Carroll, W., Silva-Alvim, F. A.L., Waghmare, S., Papanatsiou, M., Amtmann, A., Hills, A., Alvim, J. C., Blatt, M. R., Zhang, B. (2022) Engineering a K+ channel 'sensory antenna' enhances stomatal kinetics, water use efficiency and photosynthesis. Nature Plants, 8, pp. 1262-1274. (doi: 10.1038/s41477-022-01255-2)
Gracie, J., Zamberlan, F., Andrews, I. B., Smith, B. O., Peveler, W. J. (2022) Growth of plasmonic nanoparticles for aging cask-matured whisky. ACS Applied Nano Materials, 5, pp. 15362-15368. (doi: 10.1021/acsanm.2c03406)
Alharbi, A. G., Tobin, A. B., Milligan, G. (2022) How arrestins and GRKs regulate the function of long chain fatty acid receptors. International Journal of Molecular Sciences, 23, (doi: 10.3390/ijms232012237)
Ryan, S. M., Ruscher, R., Johnston, W. A., Pickering, D. A., Kennedy, M. W., Smith, B. O., Jones, L., Buitrago, G., Field, M. A., Esterman, A. J., McHugh, C. P., Browne, D. J., Cooper, M. M., Ryan, R. Y. M., Doolan, D. L., Engwerda, C. R., Miles, K., Mitreva, M., Croese, J., Rahman, T., Alexandrov, K., Giacomin, P. R., Loukas, A. (2022) Novel antiinflammatory biologics shaped by parasite–host coevolution. Proceedings of the National Academy of Sciences of the United States of America, 119, (doi: 10.1073/pnas.2202795119)
Mahindra, A., Jenkins, L., Marsango, S., Huggett, M., Huggett, M., Robinson, L., Gillespie, J., Rajamanickam, M., Morrison, A., McElroy, S., Tikhonova, I. G., Milligan, G., Jamieson, A. G. (2022) Investigating the structure–activity relationship of 1,2,4-triazine G-protein-coupled receptor 84 (GPR84) antagonists. Journal of Medicinal Chemistry, 65, pp. 11270-11290. (doi: 10.1021/acs.jmedchem.2c00804)
Kjeldsen, A., Kay, J. E., Baxter, S., McColm, S., Serrano-Amatriain, C., Parker, S., Robb, E., Arnold, S. A., Gilmour, C., Raper, A., Robertson, G., Fleming, R., Smith, B. O., Fotheringham, I. G., Christie, J. M., Magneschi, L. (2022) The fluorescent protein iLOV as a reporter for screening of high‐yield production of antimicrobial peptides in Pichia pastoris. Microbial Biotechnology, 15, pp. 2126-2139. (doi: 10.1111/1751-7915.14034)
Marsango, S., Barki, N., Jenkins, L., Tobin, A. B., Milligan, G. (2022) Therapeutic validation of an orphan G protein-coupled receptor: the case of GPR84. British Journal of Pharmacology, 179, pp. 3529-3541. (doi: 10.1111/bph.15248)
Marsango, S., Jenkins, L., Pediani, J. D., Bradley, S. J., Ward, R. J., Hesse, S., Biener, G., Stoneman, M. R., Tobin, A. B., Raicu, V., Milligan, G. (2022) The M1 muscarinic receptor is present in situ as a ligand-regulated mixture of monomers and oligomeric complexes. Proceedings of the National Academy of Sciences of the United States of America, 119, (doi: 10.1073/pnas.2201103119)
Caspar, B., Cocchiara, P., Melet, A., Van Emelen, K., Van der Aa, A., Milligan, G., Herbeuval, J.-P. (2022) CXCR4 as a novel target in immunology: moving away from typical antagonists. Future Drug Discovery, 4, (doi: 10.4155/fdd-2022-0007)
Torres Cabán, C. C., Yang, M., Lai, C., Yang, L., Subach, F. V., Smith, B. O., Piatkevich, K. D., Boyden, E. S. (2022) Tuning the sensitivity of genetically encoded fluorescent potassium indicators through structure-guided and genome mining strategies. ACS Sensors, 7, pp. 1336-1346. (doi: 10.1021/acssensors.1c02201)
Marsango, S., Ward, R. J., Jenkins, L., Butcher, A. J., Al Mahmud, Z., Dwomoh, L., Nagel, F., Schulz, S., Tikhonova, I. G., Tobin, A. B., Milligan, G. (2022) Selective phosphorylation of threonine residues defines GPR84-arrestin interactions of biased ligands. Journal of Biological Chemistry, 298, (doi: 10.1016/j.jbc.2022.101932)
Thu, N. B.A., Amtmann, A., Blatt, M. R., Nguyen, T.-H. (2022) Stomata under salt stress—what can mechanistic modeling tell us? Academic Press
Živković, D., Sanchez Dafun, A., Menneteau, T., Schahl, A., Lise, S., Kervarrec, C., Toste Rêgo, A., da Fonseca, P. C. A., Chavent, M., Pineau, C., Burlet-Schiltz, O., Marcoux, J., Bousquet, M.-P. (2022) Proteasome complexes experience profound structural and functional rearrangements throughout mammalian spermatogenesis. Proceedings of the National Academy of Sciences of the United States of America, 119, pp. e2116826119. (doi: 10.1073/pnas.2116826119)
Tibbo, A. J., Mika, D., Dobi, S., Ling, J., McFall, A., Tejeda, G. S., Blair, C., MacLeod, R., MacQuaide, N., Gök, C., Fuller, W., Smith, B. O., Smith, G. L., Vandecasteele, G., Brand, T., Baillie, G. S. (2022) Phosphodiesterase type 4 anchoring regulates cAMP signaling to Popeye domain-containing proteins. Journal of Molecular and Cellular Cardiology, 165, pp. 86-102. (doi: 10.1016/j.yjmcc.2022.01.001)
Barki, N., Bolognini, D., Börjesson, U., Jenkins, L., Riddell, J., Hughes, D. I., Ulven, T., Hudson, B. D., Rexen Ulven, E., Dekker, N., Tobin, A. B., Milligan, G. (2022) Chemogenetics defines the roles of short chain fatty acid receptors within the gut-brain axis. eLife, 11, (doi: 10.7554/eLife.73777)
Divorty, N., Jenkins, L., Ganguly, A., Butcher, A. J., Hudson, B. D., Schulz, S., Tobin, A. B., Nicklin, S. A., Milligan, G. (2022) Agonist-induced phosphorylation of orthologues of the orphan receptor GPR35 functions as an activation sensor. Journal of Biological Chemistry, 298, (doi: 10.1016/j.jbc.2022.101655)
Blatt, M. R., Jezek, M., Lew, V. L., Hills, A. (2022) What can mechanistic models tell us about guard cells, photosynthesis, and water use efficiency? Trends in Plant Science, 27, pp. 166-179. (doi: 10.1016/j.tplants.2021.08.010)
Nakasone, M. A., Majorek, K. A., Gabrielsen, M., Sibbet, G. J., Smith, B. O., Huang, D. T. (2022) Structure of UBE2K-Ub/E3/polyUb reveals mechanisms of K48-linked Ub chain extension. Nature Chemical Biology, 18, pp. 422-431. (doi: 10.1038/s41589-021-00952-x)
Neckebroeck, A., Kelly, S. M., Smith, B. O., Clark, J. S. (2022) Synthesis of the prototypical cyclopropyl dipeptide mimic and evaluation of its turn-inducing capability. Journal of Organic Chemistry, 87, pp. 258-270. (doi: 10.1021/acs.joc.1c02344)
Hudson, B. (2022) Allosteric ligands to study medium and long chain free fatty acid GPCRs. Academic Press
Zhdanov, O., Blatt, M. R., Zare-Behtash, H., Busse, A. (2022) Unidirectional versus bidirectional brushing: simulating wind influence on Arabidopsis thaliana. Quantitative Plant Biology, 3, (doi: 10.1017/qpb.2021.14)
2021
Scarpa, M., Molloy, C., Jenkins, L., Strellis, B., Budgett, R. F., Hesse, S., Dwomoh, L., Marsango, S., Tejeda, G. S., Rossi, M., Ahmed, Z., Milligan, G., Hudson, B. D., Tobin, A. B., Bradley, S. J. (2021) Biased M1 muscarinic receptor mutant mice show accelerated progression of prion neurodegenerative disease. Proceedings of the National Academy of Sciences of the United States of America, 118, pp. e2107389118. (doi: 10.1073/pnas.2107389118)
Lin, L.-C., Quon, T., Engberg, S., Mackenzie, A. E., Tobin, A. B., Milligan, G. (2021) G protein-coupled receptor GPR35 suppresses lipid accumulation in hepatocytes. ACS Pharmacology and Translational Science, 4, pp. 1835-1848. (doi: 10.1021/acsptsci.1c00224)
Blatt, M. (2021) Ae fond fareweel. Plant Physiology, 187, pp. 2341-2343. (doi: 10.1093/plphys/kiab467)
Kok, F. O., Wang, H., Riedlova, P., Goodyear, C. S., Carmody, R. J. (2021) Defining the structure of the NF-ĸB pathway in human immune cells using quantitative proteomic data. Cellular Signalling, 88, (doi: 10.1016/j.cellsig.2021.110154)
Brown, A. J.H. et al. (2021) From structure to clinic: design of a muscarinic M1 receptor agonist with potential to treatment of Alzheimer’s disease. Cell, 184, pp. 5886-5901.e22. (doi: 10.1016/j.cell.2021.11.001)
Mahindra, A., Tejeda, G., Rossi, M., Janha, O., Herbert, I., Morris, C., Morgan, D. C., Beattie, W., Montezano, A. C., Hudson, B., Tobin, A. B., Bhella, D., Touyz, R. M., Jamieson, A. G., Baillie, G. S., Blair, C. M. (2021) Peptides derived from the SARS-CoV-2 receptor binding motif bind to ACE2 but do not block ACE2-mediated host cell entry or pro-inflammatory cytokine induction. PLoS ONE, 16, (doi: 10.1371/journal.pone.0260283)
Hansen, A. H., Christensen, H. B., Pandey, S. K., Sergeev, E., Valentini, A., Dunlop, J., Dedeo, D., Fratta, S., Hudson, B., Milligan, G., Ulven, T., Ulven, E. R. (2021) Structure‐activity relationship explorations and discovery of a potent antagonist for the free fatty acid receptor 2. ChemMedChem, 16, (doi: 10.1002/cmdc.202100356)
Somma, D., Kok, F. O., Kerrigan, D., Wells, C. A., Carmody, R. J. (2021) Defining the role of nuclear factor NF-κB p105 subunit in human macrophage by transcriptomic analysis of NFKB1 knockout THP-1 cells. Frontiers in Immunology, 12, (doi: 10.3389/fimmu.2021.669906)
Milligan, G., Jenkins, L., Marsango, S., Mancini, S., Mahmud, Z. A., Morrison, A., McElroy, S., Bennett, K., Barnes, M., Tobin, A., Tikhonova, I. (2021) Discovery and characterization of novel antagonists of the proinflammatory orphan receptor GPR84. ACS Pharmacology and Translational Science, 4, pp. 1598-1613. (doi: 10.1021/acsptsci.1c00151)
Prabhakar, A. T., James, C. D., Das, D., Otoa, R., Day, M., Burgner, J., Fontan, C. T., Wang, X., Wieland, A., Donaldson, M. M., Bristol, M. L., Li, R., Oliver, A. W., Pearl, L. H., Smith, B. O., Glass, S., Morgan, I. M. (2021) CK2 phosphorylation of human papillomavirus 16 E2 on serine 23 promotes interaction with TopBP1 and is critical for E2 interaction with mitotic chromatin and the viral life cycle. mBio, 12, (doi: 10.1128/mbio.01163-21)
Buist, H. K., Luchowska-Stańska, U., van Basten, B., Valli, J., Smith, B. O., Baillie, G. S., Rickman, C., Ricketts, B., Davidson, A., Hannam, R., Sunderland, J., Yarwood, S. J. (2021) Identification and characterization of an affimer affinity reagent for the detection of the cAMP sensor, EPAC1. Cells, 10, (doi: 10.3390/cells10092307)
Ahwazi, D., Neopane, K., Markby, G. R., Kopietz, F., Ovens, A. J., Dall, M., Hassing, A. S., Graesle, P., Alshuweishi, Y. A. I., Treebak, J. T., Salt, I., Göransson, O., Zeqiraj, E., Scott, J. W., Sakamoto, K. (2021) Investigation of the specificity and mechanism of action of the ULK1/AMPK inhibitor SBI-0206965. Biochemical Journal, 478, pp. 2977-2997. (doi: 10.1042/BCJ20210284)
Palmer, T. M., Salt, I. P. (2021) Nutrient regulation of inflammatory signalling in obesity and vascular disease. Clinical Science, 135, pp. 1563-1590. (doi: 10.1042/CS20190768)
Morgan, D. C., Morris, C., Mahindra, A., Blair, C. M., Tejeda, G., Herbert, I., Turnbull, M. L., Lieber, G., Willett, B. J., Logan, N., Smith, B., Tobin, A. B., Bhella, D., Baillie, G. S., Jamieson, A. G. (2021) Stapled ACE2 peptidomimetics designed to target the SARS-CoV-2 spike protein do not prevent virus internalisation. Peptide Science, 113, (doi: 10.1002/pep2.24217)
Jezek, M., Silva-Alvim, F. A.L., Hills, A., Donald, N., Ishka, M. R., Shadbolt, J., He, B., Lawson, T., Harper, J. F., Wang, Y., Lew, V. L., Blatt, M. R. (2021) Guard cell endomembrane Ca2+-ATPases underpin a ‘carbon memory’ of photosynthetic assimilation that impacts on water-use efficiency. Nature Plants, 7, pp. 1301-1313. (doi: 10.1038/s41477-021-00966-2)
Boleij, A., Fathi, P., Dalton, W., Park, B., Wu, X., Huso, D., Allen, J., Besharati, S., Anders, R. A., Housseau, F., Mackenzie, A. E., Jenkins, L., Milligan, G., Wu, S., Sears, C. L. (2021) G-protein coupled receptor 35 (GPR35) regulates the colonic epithelial cell response to enterotoxigenic Bacteroides fragilis. Communications Biology, 4, (doi: 10.1038/s42003-021-02014-3)
Salt, I. P., Nunes, J. R.C., Fullerton, M. D. (2021) Metformin again? Atheroprotection mediated by macrophage AMPK and ATF1. Cardiovascular Research, 117, pp. 1233-1234. (doi: 10.1093/cvr/cvab065)
Morris, E. P., da Fonseca, P. C.A. (2021) How to build a proteasome. Nature Structural and Molecular Biology, 28, pp. 409-410. (doi: 10.1038/s41594-021-00592-8)
Klejchova, M., Silva-Alvim, F. A.L., Blatt, M. R., Alvim, J. C. (2021) Membrane voltage as a dynamic platform for spatio-temporal signalling, physiological and developmental regulation. Plant Physiology, 185, pp. 1523-1541. (doi: 10.1093/plphys/kiab032)
Milligan, G., Barki, N., Tobin, A. (2021) Chemogenetic approaches to explore the functions of Free Fatty Acid Receptor 2. Trends in Pharmacological Sciences, 42, pp. 191-202. (doi: 10.1016/j.tips.2020.12.003)
Zhdanov, O., Blatt, M. R., Zare-Behtash, H., Busse, A. (2021) Wind-evoked anemotropism affects the morphology and mechanical properties of Arabidopsis. Journal of Experimental Botany, 72, pp. 1906-1918. (doi: 10.1093/jxb/eraa541)
Kopietz, F., Alshuweishi, Y., Bijland, S., Alghamdi, F., Degerman, E., Sakamoto, K., Salt, I. P., Göransson, O. (2021) A-769662 inhibits adipocyte glucose uptake in an AMPK-independent manner. Biochemical Journal, 478, pp. 633-646. (doi: 10.1042/BCJ20200659)
Bolognini, D., Dedeo, D., Milligan, G. (2021) Metabolic and inflammatory functions of short-chain fatty acid receptors. Current Opinion in Endocrine and Metabolic Research, 16, pp. 1-9. (doi: 10.1016/j.coemr.2020.06.005)
Wong, J. H., Klejchova, M., Snipes, S. A., Nagpal, P., Bak, G., Wang, B., Dunlap, S., Park, M. Y., Kunkel, E. N., Trinidad, B., Reed, J. W., Blatt, M. R., Gray, W. M. (2021) SAUR proteins and PP2C.D phosphatases regulate H+-ATPases and K+ channels to control stomatal movements. Plant Physiology, 185, pp. 256-273. (doi: 10.1093/plphys/kiaa023)
Knuhtsen, A., Whiting, R., McWhinnie, F. S., Whitmore, C., Smith, B. O., Green, A. C., Timperley, C. M., Kinnear, K. I., Jamieson, A. G. (2021) μ-conotoxin KIIIA peptidomimetics that block human voltage-gated sodium channels. Peptide Science, 113, (doi: 10.1002/pep2.24203)
Ward, R. J., Pediani, J. D., Marsango, S., Jolly, R., Stoneman, M. R., Biener, G., Handel, T. M., Raicu, V., Milligan, G. (2021) Chemokine receptor CXCR4 oligomerization is disrupted selectively by the antagonist ligand IT1t. Journal of Biological Chemistry, 296, (doi: 10.1074/jbc.RA120.016612)
2020
Zhdanov, O., Blatt, M. R., Cammarano, A., Zare-Behtash, H., Busse, A. (2020) A new perspective on mechanical characterisation of Arabidopsis stems through vibration tests. Journal of the Mechanical Behavior of Biomedical Materials, 112, (doi: 10.1016/j.jmbbm.2020.104041)
Alghamdi, F., Alshuweishi, Y., Salt, I. P. (2020) Regulation of nutrient uptake by AMP-activated protein kinase. Cellular Signalling, 76, (doi: 10.1016/j.cellsig.2020.109807)
Marti-Solano, M., Crilly, S. E., Malinverni, D., Munk, C., Harris, M., Pearce, A., Quon, T., Mackenzie, A. E., Wang, X., Peng, J., Tobin, A. B., Ladds, G., Milligan, G., Gloriam, D. E., Puthenveedu, M. A., Babu, M. M. (2020) Combinatorial expression of GPCR isoforms affects signalling and drug responses. Nature, 587, pp. 650-656. (doi: 10.1038/s41586-020-2888-2)
Labéguère, F. et al. (2020) Discovery of 9-Cyclopropylethynyl-2-((S)-1-[1,4]dioxan-2-ylmethoxy)-6,7-dihydropyrimido[6,1-a]isoquinolin-4-one (GLPG1205), a unique GPR84 negative allosteric modulator undergoing evaluation in a phase II clinical trial. Journal of Medicinal Chemistry, 63, pp. 13526-13545. (doi: 10.1021/acs.jmedchem.0c00272)
Chatrin, C., Gabrielsen, M., Buetow, L., Nakasone, M. A., Ahmed, S. F., Sumpton, D., Sibbet, G. J., Smith, B. O., Huang, D. T. (2020) Structural insights into ADP-ribosylation of ubiquitin by Deltex family E3 ubiquitin ligases. Science Advances, 6, (doi: 10.1126/sciadv.abc0418)
Lefoulon, C., Boxall, S. F., Hartwell, J., Blatt, M. R. (2020) Crassulacean acid metabolism guard cell anion channel activity follows transcript abundance and is suppressed by apoplastic malate. New Phytologist, 227, pp. 1847-1857. (doi: 10.1111/nph.16640)
Larson, E. R., Ortmannová, J., Donald, N. A., Alvim, J., Blatt, M. R., Žárský, V. (2020) Synergy among exocyst and SNARE interactions identifies a functional hierarchy in secretion during vegetative growth. Plant Cell, 32, pp. 2951-2963. (doi: 10.1105/tpc.20.00280)
Prihandoko, R., Kaur, D., Wiegman, C. H., Alvarez-Curto, E., Donovan, C., Chachi, L., Ulven, T., Tyas, M. R., Euston, E., Dong, Z., Alharbi, A. G. M., Kim, R. Y., Lowe, J. G., Hansbro, P. M., Chung, K. F., Brightling, C. E., Milligan, G., Tobin, A. B. (2020) Pathophysiological regulation of lung function by the free fatty acid receptor FFA4. Science Translational Medicine, 12, (doi: 10.1126/scitranslmed.aaw9009)
Mitxitorena, I., Somma, D., Mitchell, J. P., Lepistö, M., Tyrchan, C., Smith, E. L., Kiely, P. A., Walden, H., Keeshan, K., Carmody, R. J. (2020) The deubiquitinase USP7 uses a distinct ubiquitin-like domain to deubiquitinate NF-ĸB subunits. Journal of Biological Chemistry, 295, pp. 11754-11763. (doi: 10.1074/jbc.RA120.014113)
Jezek, M., Hills, A., Blatt, M. R., Lew, V. L. (2020) A constraint-relaxation-recovery mechanism for stomatal dynamics. Plant, Cell and Environment, 42, pp. 2399-2410. (doi: 10.1111/pce.13568)
Quon, T., Lin, L.-C., Ganguly, A., Tobin, A. B., Milligan, G. (2020) Therapeutic opportunities and challenges in targeting the orphan G protein-coupled receptor GPR35. ACS Pharmacology and Translational Science, 3, pp. 801-812. (doi: 10.1021/acsptsci.0c00079)
Flütsch, S., Wang, Y., Takemiya, A., Vialet-Chabrand, S. R., Klejchova, M., Nigro, A., Hills, A., Lawson, T., Blatt, M. R., Santelia, D. (2020) Guard cell starch degradation yields glucose for rapid stomatal opening in Arabidopsis. Plant Cell, 32, pp. 2325-2344. (doi: 10.1105/tpc.18.00802)
Sarrou, E., Richmond, L., Carmody, R. J., Gibson, B., Keeshan, K. (2020) CRISPR gene editing of murine blood stem and progenitor cells induces MLL-AF9 chromosomal translocation and MLL-AF9 leukaemogenesis. International Journal of Molecular Sciences, 21, (doi: 10.3390/ijms21124266)
Klejchova, M., Hills, A., Blatt, M. R. (2020) Predicting the unexpected in stomatal gas exchange: not just an open-and-shut case. Biochemical Society Transactions, 48, pp. 881-889. (doi: 10.1042/BST20190632)
Crecente Garcia, S., Neckebroeck, A., Clark, J. S., Smith, B. O., Thomson, A. R. (2020) β-turn mimics by chemical ligation. Organic Letters, 22, pp. 4424-4428. (doi: 10.1021/acs.orglett.0c01427)
Rexen Ulven, E., Quon, T., Sergeev, E., Barki, N., Brvar, M., Hudson, B. D., Dutta, P., Højgaard Hansen, A., Bielefeldt, L. Ø., Tobin, A. B., McKenzie, C. J., Milligan, G., Ulven, T. (2020) Structure-activity relationship studies of tetrahydroquinolone free fatty acid receptor 3 modulators. Journal of Medicinal Chemistry, 63, pp. 3577-3595. (doi: 10.1021/acs.jmedchem.9b02036)
Horaruang, W., Hills, A., Blatt, M. R. (2020) Communication between the plasma membrane and tonoplast is an emergent property of ion transport. Plant Physiology, 182, pp. 1833-1835. (doi: 10.1104/pp.19.01485)
Sencio, V., Barthelemy, A., Tavares, L. P., Machado, M. G., Soulard, D., Cuinat, C., Queiroz-Junior, C. M., Noordine, M. L., Salomé-Desnoulez, S., Deryuter, L., Foligné, B., Wahl, C., Frisch, B., Vieira, A. T., Paget, C., Milligan, G., Ulven, T., Wolowczuk, I., Faveeuw, C., Le Goffic, R., Thomas, M., Ferreira, S., Teixeira, M. M., Trottein, F. (2020) Gut dysbiosis during influenza contributes to pulmonary pneumococcal superinfection through altered short-chain fatty acid production. Cell Reports, 30, pp. 2934-2947.e6. (doi: 10.1016/j.celrep.2020.02.013)
2019
Collins, P. E., Somma, D., Kerrigan, D., Herrington, F., Keeshan, K. R., Nibbs, R. J.B., Carmody, R. J. (2019) The IκB-protein BCL-3 controls toll-like receptor-induced MAPK activity by promoting TPL-2 degradation in the nucleus. Proceedings of the National Academy of Sciences of the United States of America, 116, pp. 25828-25838. (doi: 10.1073/pnas.1900408116)
Heuninck, J., Perpina Viciano, C., Işbilir, A., Caspar, B., Capoferri, D., Briddon, S. J., Durroux, T., Hill, S. J., Lohse, M. J., Milligan, G., Pin, J.-P., Hoffmann, C. (2019) Context-dependent signaling of CXC chemokine receptor 4 and atypical chemokine receptor 3. Molecular Pharmacology, 96, pp. 778-793. (doi: 10.1124/mol.118.115477)
Smith, E. L., Somma, D., Kerrigan, D., McIntyre, Z., Cole, J. J., Liang, K. L., Kiely, P. A., Keeshan, K., Carmody, R. J. (2019) The regulation of sequence specific NF-ĸB DNA binding and transcription by IKKβ Phosphorylation of NF-ĸB p50 at Serine 80. Nucleic Acids Research, 47, pp. 11151-11161. (doi: 10.1093/nar/gkz873)
Waghmare, S., Lefoulon, C., Zhang, B., Lileikyte, E., Donald, N., Blatt, M. (2019) K+ channel and SEC11 binding exchange regulates SNARE assembly for secretory traffic. Plant Physiology, 181, pp. 1096-1113. (doi: 10.1104/pp.19.00919)
Zhdanov, O., Busse, A., Cammarano, A., Zare-Behtash, H., Blatt, M. (2019) Effect of Wind on Morphology and Mechanical Properties of Arabidopsis Thaliana.
Toste Rêgo, A., da Fonseca, P. C.A. (2019) Characterization of fully recombinant human 20S and 20S-PA200 proteasome complexes. Molecular Cell, 76, pp. 138-147.e5. (doi: 10.1016/j.molcel.2019.07.014)
Jurić, I., Hibberd, J. M., Blatt, M., Burroughs, N. J. (2019) Computational modelling predicts substantial carbon assimilation gains for C3 plants with a single-celled C4 biochemical pump. PLoS Computational Biology, 15, (doi: 10.1371/journal.pcbi.1007373)
Ibáñez-Shimabukuro, M., Rey-Burusco, M. F., Gabrielsen, M., Franchini, G. R., Riboldi-Tunnicliffe, A., Roe, A. J., Griffiths, K., Cooper, A., Córsico, B., Kennedy, M. W., Smith, B. O. (2019) Structure and ligand binding of As-p18, an extracellular fatty acid binding protein from the eggs of a parasitic nematode. Bioscience Reports, 39, (doi: 10.1042/BSR20191292)
Katwan, O. J., Alghamdi, F., Almabrouk, T. A., Mancini, S. J., Kennedy, S., Oakhill, J. S., Scott, J. W., Salt, I. P. (2019) AMP-activated protein kinase complexes containing the β2 regulatory subunit are upregulated during and contribute to adipogenesis. Biochemical Journal, 476, pp. 1725-1740. (doi: 10.1042/BCJ20180714)
Blackman, M. J., Stokes, B. H., Yoo, E., Murithi, J. M., Luth, M. R., Afanasyev, P., da Fonseca, P. C.A., Winzeler, E. A., Ng, C. L., Bogyo, M., Fidock, D. A. (2019) Covalent Plasmodium falciparum-selective proteasome inhibitors exhibit a low propensity for generating resistance in vitro and synergize with multiple antimalarial agents. PLoS Pathogens, 15, (doi: 10.1371/journal.ppat.1007722)
Stoneman, M. R., Biener, G., Ward, R. J., Pediani, J. D., Badu, D., Eis, A., Popa, I., Milligan, G., Raicu, V. (2019) A general method to quantify ligand-driven oligomerization from fluorescence-based images. Nature Methods, 16, pp. 493-496. (doi: 10.1038/s41592-019-0408-9)
Bolognini, D., Barki, N., Butcher, A. J., Hudson, B. D., Sergeev, E., Molloy, C., Moss, C. E., Bradley, S. J., Le Gouill, C., Bouvier, M., Tobin, A. B., Milligan, G. (2019) Chemogenetics defines receptor-mediated functions of short chain free fatty acids. Nature Chemical Biology, 15, pp. 489-498. (doi: 10.1038/s41589-019-0270-1)
Zhang, B., Karnik, R., Alvim, J., Donald, N., Blatt, M. R. (2019) Dual sites for SEC11 on the SNARE SYP121 implicate a binding exchange during secretory traffic. Plant Physiology, 180, pp. 228-239. (doi: 10.1104/pp.18.01315)
Milligan, G., Ward, R. J., Marsango, S. (2019) GPCR homo-oligomerization. Current Opinion in Cell Biology, 57, pp. 40-47. (doi: 10.1016/j.ceb.2018.10.007)
Mackenzie, A. E., Quon, T., Lin, L.-C., Hauser, A. S., Jenkins, L., Inoue, A., Tobin, A. B., Gloriam, D. E., Hudson, B. D., Milligan, G. (2019) Receptor selectivity between the G proteins Gα12 and Gα13 is defined by a single leucine-to-isoleucine variation. FASEB Journal, 33, pp. 5005-5017. (doi: 10.1096/fj.201801956R)
Papanatsiou, M., Petersen, J., Henderson, L., Wang, Y., Christie, J.M., Blatt, M.R. (2019) Optogenetic manipulation of stomatal kinetics improves carbon assimilation, water use, and growth. Science, 363, pp. 1456-1459. (doi: 10.1126/science.aaw0046)
Zhao, C. et al. (2019) Evolution of chloroplast retrograde signaling facilitates green plant adaptation to land. Proceedings of the National Academy of Sciences of the United States of America, 116, pp. 5015-5020. (doi: 10.1073/pnas.1812092116)
Knuhtsen, A., Whitmore, C., McWhinnie, F. S., McDougall, L., Whiting, R., Smith, B. O., Timperley, C. M., Green, A. C., Kinnear, K. I., Jamieson, A. G. (2019) α-conotoxin GI triazole-peptidomimetics: potent and stable blockers of a human acetylcholine receptor. Chemical Science, 10, pp. 1671-1676. (doi: 10.1039/C8SC04198A)
Mancini, S., Mahmud, Z. A., Jenkins, L., Bolognini, D., Newman, R., Barnes, M., Edye, M. E., McMahon, S. B., Tobin, A. B., Milligan, G. (2019) On-target and off-target effects of novel orthosteric and allosteric activators of GPR84. Scientific Reports, 9, (doi: 10.1038/s41598-019-38539-1)
Liao, X., Zhang, B., Blatt, M. R., Jenkins, G. I. (2019) A FRET method for investigating dimer/monomer status and conformation of the UVR8 photoreceptor. Photochemical and Photobiological Sciences, 18, pp. 367-374. (doi: 10.1039/C8PP00489G)
Müskens, F. M., Ward, R. J., Herkt, D., van de Langemheen, H., Tobin, A. B., Liskamp, R. M.J., Milligan, G. (2019) Design, synthesis and evaluation of a diazirine photoaffinity probe for ligand-based receptor capture targeting G protein-coupled receptors. Molecular Pharmacology, 95, pp. 196-209. (doi: 10.1124/mol.118.114249)
Petrie, J. R., Salt, I. P. (2019) Diabetes and vascular disease. Springer
Rezig, I. M., Bremner, S. K., Bhutta, M. S., Salt, I. P., Gould, G. W., McInerny, C. J. (2019) Genetic and cytological methods to study ESCRT cell cycle function in fission yeast. Humana Press
2018
Strembitska, A., Mancini, S. J., Gamwell, J. M., Palmer, T. M., Baillie, G. S., Salt, I. P. (2018) A769662 inhibits insulin-stimulated Akt activation in human macrovascular endothelial cells independent of AMP-activated protein kinase. International Journal of Molecular Sciences, 19, (doi: 10.3390/ijms19123886)
Waghmare, S., Lileikyte, E., Karnik, R., Goodman, J. K., Blatt, M. R., Jones, A. M.E. (2018) SNAREs SYP121 and SYP122 mediate the secretion of distinct cargo subsets. Plant Physiology, 178, pp. 1679-1688. (doi: 10.1104/pp.18.00832)
Kennedy, A. J., Ballante, F., Johansson, J. R., Milligan, G., Sunström, L., Nordqvist, A., Carlsson, J. (2018) Structural characterization of agonist binding to protease-activated receptor 2 through mutagenesis and computational modeling. ACS Pharmacology and Translational Science, 1, pp. 119-133. (doi: 10.1021/acsptsci.8b00019)
Hansen, A. H., Sergeev, E., Bolognini, D., Sprenger, R. R., Ekberg, J. H., Ejsing, C. S., McKenzie, C. J., Rexen Ulven, E., Milligan, G., Ulven, T. (2018) Discovery of a potent thiazolidine free fatty acid receptor 2 agonist with favorable pharmacokinetic properties. Journal of Medicinal Chemistry, 61, pp. 9534-9550. (doi: 10.1021/acs.jmedchem.8b00855)
Lefoulon, C., Waghmare, S., Karnik, R., Blatt, M. R. (2018) Gating control and K+ uptake by the KAT1 K+ channel leaveraged through membrane anchoring of the trafficking protein SYP121. Plant, Cell and Environment, 41, pp. 2668-2677. (doi: 10.1111/pce.13392)
Drew, J. E., Reichardt, N., Williams, L. M., Mayer, C.-D., Walker, A. W., Farquharson, A. J., Kastora, S., Farquharson, F., Milligan, G., Morrison, D. J., Preston, T., Flint, H. J., Louis, P. (2018) Dietary fibers inhibit obesity in mice, but host responses in the cecum and liver appear unrelated to fiber-specific changes in cecal taxonomic composition. Scientific Reports, 8, (doi: 10.1038/s41598-018-34081-8)
Zhang, B., Karnik, R., Donald, N., Blatt, M. R. (2018) A GPI signal peptide-anchored split-ubiquitin (GPS) system for detecting soluble bait protein interactions at the membrane. Plant Physiology, 178, pp. 13-17. (doi: 10.1104/pp.18.00577)
Divorty, N., Milligan, G., Graham, D., Nicklin, S. A. (2018) The orphan receptor GPR35 contributes to angiotensin II–induced hypertension and cardiac dysfunction in mice. American Journal of Hypertension, 31, pp. 1049-1058. (doi: 10.1093/ajh/hpy073)
Feroz, H., Ferlez, B., Lefoulon, C., Ren, T., Baker, C. S., Gajewski, J. P., Lugar, D. J., Gaudana, S. B., Butler, P. J., Hühn, J., Lamping, M., Parak, W. J., Hibberd, J. M., Kerfeld, C. A., Smirnoff, N., Blatt, M. R., Golbeck, J. H., Kumar, M. (2018) Light-driven chloride transport kinetics of halorhodopsin. Biophysical Journal, 115, pp. 353-360. (doi: 10.1016/j.bpj.2018.06.009)
Marsango, S., Ward, R. J., Alvarez-Curto, E., Milligan, G. (2018) Muscarinic receptor oligomerization. Neuropharmacology, 136, pp. 401-410. (doi: 10.1016/j.neuropharm.2017.11.023)
Binti Mohd Amir, N. A.S., Mackenzie, A. E., Jenkins, L., Boustani, K., Hillier, M. C., Tsuchiya, T., Milligan, G., Pease, J. E. (2018) Evidence for the existence of a CXCL17 receptor distinct from GPR35. Journal of Immunology, 201, pp. 714-724. (doi: 10.4049/jimmunol.1700884)
Milligan, G. (2018) G protein-coupled receptors not currently in the spotlight: free fatty acid receptor 2 and GPR35. British Journal of Pharmacology, 175, pp. 2543-2553. (doi: 10.1111/bph.14042)
Zhdanov, O., Busse, A., Blatt, M., Zare-Behtash, H. (2018) A specialised wind tunnel for investigation of ecophysiological response of plants to wind.
Butcher, S. K., O'Carroll, C. E., Wells, C. A., Carmody, R. J. (2018) Toll-like receptors drive specific patterns of tolerance and training on restimulation of macrophages. Frontiers in Immunology, 9, (doi: 10.3389/fimmu.2018.00933)
Salomé, M., Magee, A., Yalla, K., Chaudhury, S., Sarrou, E., Carmody, R. J., Keeshan, K. (2018) A Trib2-p38 axis controls myeloid leukaemia cell cycle and stress response signalling. Cell Death and Disease, 9, (doi: 10.1038/s41419-018-0467-3)
Milligan, G., Inoue, A. (2018) Genome editing provides new insights into receptor-controlled signalling pathways. Trends in Pharmacological Sciences, 39, pp. 481-493. (doi: 10.1016/j.tips.2018.02.005)
Blatt, M. R. (2018) Concepts and techniques in plant membrane physiology. John Wiley & Sons, Ltd
Blatt, M.R., Thiel, G. (2018) SNARE components and mechanisms of exocytosis in plants. John Wiley & Sons, Ltd
Mancini, S. J., Boyd, D., Katwan, O. J., Strembitska, A., Almabrouk, T. A., Kennedy, S., Palmer, T. M., Salt, I. P. (2018) Canagliflozin inhibits interleukin-1β-stimulated cytokine and chemokine secretion in vascular endothelial cells by AMP-activated protein kinase-dependent and -independent mechanisms. Scientific Reports, 8, (doi: 10.1038/s41598-018-23420-4)
Almabrouk, T. A.M., White, A. D., Ugusman, A. B., Skiba, D. S., Katwan, O. J., Alganga, H., Guzik, T. J., Touyz, R. M., Salt, I. P., Kennedy, S. (2018) High fat diet attenuates the anticontractile activity of aortic PVAT via a mechanism involving AMPK and reduced adiponectin secretion. Frontiers in Physiology, 9, (doi: 10.3389/fphys.2018.00051)
Speirs, C., Williams, J. J.L., Riches, K., Salt, I. P., Palmer, T. M. (2018) Linking energy sensing to suppression of JAK-STAT signalling: a potential route for repurposing AMPK activators? Pharmacological Research, 128, pp. 88-100. (doi: 10.1016/j.phrs.2017.10.001)
Pediani, J. D., Ward, R. J., Marsango, S., Milligan, G. (2018) Spatial intensity distribution analysis: studies of G Protein-coupled receptor oligomerization. Trends in Pharmacological Sciences, 39, pp. 175-186. (doi: 10.1016/j.tips.2017.09.001)
Pantin, F., Blatt, M. R. (2018) Stomatal response to humidity: blurring the boundary between active and passive movement. Plant Physiology, 176, pp. 485-488. (doi: 10.1104/pp.17.01699)
Mancini, S. J., Salt, I. P. (2018) Investigating the role of AMPK in inflammation. Humana Press
Mitchell, J. P., Carmody, R. J. (2018) NF-κB and the Transcriptional Control of Inflammation. Elsevier
Reichardt, N., Vollmer, M., Holtrop, G., Farquharson, F. M., Wefers, D., Bunzel, M., Duncan, S. H., Drew, J. E., Williams, L. M., Milligan, G., Preston, T., Morrison, D., Flint, H. J., Louis, P. (2018) Specific substrate-driven changes in human faecal microbiota composition contrast with functional redundancy in short-chain fatty acid production. ISME Journal, 12, pp. 610-622. (doi: 10.1038/ismej.2017.196)
2017
Mahmud, Z. A., Jenkins, L., Ulven, T., Labéguère, F., Gosmini, R., De Vos, S., Hudson, B. D., Tikhonova, I. G., Milligan, G. (2017) Three classes of ligands each bind to distinct sites on the orphan G protein-coupled receptor GPR84. Scientific Reports, 7, (doi: 10.1038/s41598-017-18159-3)
Viskaitis, P., Irvine, E. E., Smith, M. A., Choudhury, A. I., Alvarez-Curto, E., Glegola, J. A., Hardy, D. G., Pedroni, S. M.A., Paiva Pessoa, M. R., Fernando, A. B.P., Katsouri, L., Sardini, A., Ungless, M. A., Milligan, G., Withters, D. J. (2017) Modulation of SF1 neuron activity coordinately regulates both feeding behaviour and associated emotional states. Cell Reports, 21, pp. 3559-3572. (doi: 10.1016/j.celrep.2017.11.089)
Cooper, A., Vance, S. J., Smith, B. O., Kennedy, M. W. (2017) Frog foams and natural protein surfactants. Colloids and Surfaces A: Physicochemical and Engineering Aspects, 534, pp. 120-129. (doi: 10.1016/j.colsurfa.2017.01.049)
Wang, Y., Hills, A., Vialet-Chabrand, S., Papanatsiou, M., Griffiths, H., Rogers, S., Lawson, T., Lew, V. L., Blatt, M. R. (2017) Unexpected connections between humidity and ion transport discovered using a model to bridge guard cell-to-leaf scales. Plant Cell, 29, pp. 2921-2939. (doi: 10.1105/tpc.17.00694)
Sergeev, E., Hansen, A. H., Bolognini, D., Kawakami, K., Kishi, T., Aoki, J., Ulven, T., Inoue, A., Hudson, B. D., Milligan, G. (2017) A single extracellular amino acid in Free Fatty Acid Receptor 2 defines antagonist species selectivity and G protein selection bias. Scientific Reports, 7, (doi: 10.1038/s41598-017-14096-3)
Gabrielsen, M., Buetow, L., Nakasone, M. A., Ahmed, S. F., Sibbet, G. J., Smith, B. O., Zhang, W., Sidhu, S. S., Huang, D. T. (2017) A general strategy for discovery of inhibitors and activators of RING and U-box E3 ligases with ubiquitin variants. Molecular Cell, 68, pp. 456-470.e10. (doi: 10.1016/j.molcel.2017.09.027)
Larson, E. R., Van Zelm, E., Roux, C., Marion-Poll, A., Blatt, M. R. (2017) Clathrin Heavy Chain subunits coordinate endo- and exocytic traffic and affect stomatal movement. Plant Physiology, 175, pp. 708-720. (doi: 10.1104/pp.17.00970)
Almabrouk, T. A. M., Ugusman, A. B., Katwan, O. J., Salt, I. P., Kennedy, S. (2017) Deletion of AMPKα1 attenuates the anticontractile effect of perivascular adipose tissue (PVAT) and reduces adiponectin release. British Journal of Pharmacology, 174, pp. 3398-3410. (doi: 10.1111/bph.13633)
Kennedy, S., Salt, I. P. (2017) Molecular mechanisms regulating perivascular adipose tissue - potential pharmacological targets? British Journal of Pharmacology, 174, pp. 3385-3387. (doi: 10.1111/bph.13969)
Magennis, S., Toulmin, A., Baltierra-Jasso, L. E., Morten, M. J., Sabir, T., McGlynn, P., Schröder, G. F., Smith, B. O., Magennis, S. W. (2017) Conformational heterogeneity in a fully-complementary DNA three-way junction with a GC-rich branchpoint. Biochemistry, 56, pp. 4985-4991. (doi: 10.1021/acs.biochem.7b00677)
Milligan, G., Alvarez-Curto, E., Hudson, B. D., Prihandoko, R., Tobin, A. B. (2017) FFA4/GPR120: pharmacology and therapeutic opportunities. Trends in Pharmacological Sciences, 38, pp. 809-821. (doi: 10.1016/j.tips.2017.06.006)
(2017) G-Protein-Coupled Receptor Dimers. (doi: 10.1007/978-3-319-60174-8)
Ward, R. J., Marsango, S., Pediani, J. D., Milligan, G. (2017) The use of spatial intensity distribution analysis to examine G protein-coupled receptor oligomerization. Humana Press
Prokop, S., Perry, N. A., Vishnivetskiy, S. A., Toth, A. D., Inoue, A., Milligan, G., Iverson, T. M., Hunyady, L., Gurevich, V. V. (2017) Differential manipulation of arrestin-3 binding to basal and agonist-activated G protein-coupled receptors. Cellular Signalling, 36, pp. 98-107. (doi: 10.1016/j.cellsig.2017.04.021)
Spies, M., Smith, B. O. (2017) Protein-nucleic acids interactions: new ways of connecting structure, dynamics and function. Biophysical Reviews, 9, pp. 289-291. (doi: 10.1007/s12551-017-0284-4)
Lopez-Gimenez, J., Alvarez-Curto, E., Milligan, G. (2017) M3 muscarinic acetylcholine receptor facilitates the endocytosis of mu opioid receptor mediated by morphine independently of the formation of heteromeric complexes. Cellular Signalling, 35, pp. 208-222. (doi: 10.1016/j.cellsig.2017.04.006)
Cai, S., Papanatsiou, M., Blatt, M. R., Chen, Z.-H. (2017) Speedy grass stomata: emerging molecular and evolutionary features. Molecular Plant, 10, pp. 912-914. (doi: 10.1016/j.molp.2017.06.002)
Riedelsberger, J., Blatt, M. R. (2017) Editorial: Roots—the hidden provider. Frontiers in Plant Science, 8, (doi: 10.3389/fpls.2017.01021)
Shaw, A., Jeromson, S., Watterson, K. R., Pediani, J. D., Gallagher, I., Whalley, T., Dreczkowski, G., Brooks, N., Galloway, S., Hamilton, D. L. (2017) Multiple AMPK activators inhibit L-Carnitine uptake in C2C12 skeletal muscle myotubes. American Journal of Physiology: Cell Physiology, 312, pp. C689-C696. (doi: 10.1152/ajpcell.00026.2016)
Cai, S., Chen, G., Wang, Y., Huang, Y., Marchant, B., Wang, Y., Yang, Q., Dai, F., Hills, A., Franks, P. J., Nevo, E., Soltis, D., Soltis, P., Sessa, E., Wolf, P. G., Xue, D., Zhang, G., Pogson, B. J., Blatt, M. R., Chen, Z.-H. (2017) Evolutionary conservation of ABA signaling for stomatal closure in ferns. Plant Physiology, 174, pp. 732-747. (doi: 10.1104/pp.16.01848)
Vialet-Chabrand, S., Hills, A., Wang, Y., Griffiths, H., Lew, V. L., Lawson, T., Blatt, M. R., Rogers, S. (2017) Global sensitivity analysis of OnGuard models identifies key hubs for transport interaction in stomatal dynamics. Plant Physiology, 174, pp. 680-688. (doi: 10.1104/pp.17.00170)
Morris, E. P., da Fonseca, P. C.A. (2017) High-resolution cryo-EM proteasome structures in drug development. Acta Crystallographica. Section D: Structural Biology, 73, pp. 522-533. (doi: 10.1107/S2059798317007021)
Watterson, K. R., Hansen, S. V.F., Hudson, B., Alvarez-Curto, E., Raihan, S. Z., Azevedo, C. M.G., Martin, G., Dunlop, J., Yarwood, S. J., Ulven, T., Milligan, G. (2017) Probe-dependent negative allosteric modulators of the long-chain free fatty acid receptor FFA4. Molecular Pharmacology, 91, pp. 630-641. (doi: 10.1124/mol.116.107821)
Ward, R. J., Pediani, J. D., Harikumar, K. G., Miller, L. J., Milligan, G. (2017) Spatial intensity distribution analysis quantifies the extent and regulation of homodimerization of the secretin receptor. Biochemical Journal, 474, pp. 1879-1895. (doi: 10.1042/BCJ20170184)
Kaspersen, M. H., Jenkins, L., Dunlop, J., Milligan, G., Ulven, T. (2017) Succinct synthesis of saturated hydroxy fatty acids and in vitro evaluation of all hydroxylauric acids on FFA1, FFA4 and GPR84. MedChemComm, 8, pp. 1360-1365. (doi: 10.1039/c7md00130d)
Vialet-Chabrand, S. R.M., Matthews, J. S.A., McAusland, L., Blatt, M. R., Griffiths, H., Lawson, T. (2017) Temporal dynamics of stomatal behavior: modeling and implications for photosynthesis and water use. Plant Physiology, 174, pp. 603-613. (doi: 10.1104/pp.17.00125)
Jezek, M., Blatt, M. R. (2017) The membrane transport system of the guard cell and its integration for stomatal dynamics. Plant Physiology, 174, pp. 487-519. (doi: 10.1104/pp.16.01949)
Salt, I. P., Hardie, G. (2017) AMP-activated protein kinase: an ubiquitous signalling pathway with key roles in the cardiovascular system. Circulation Research, 120, pp. 1825-1841. (doi: 10.1161/CIRCRESAHA.117.309633)
Marsango, S., Caltabiano, G., Jiménez-Rosés, M., Millan, M. J., Pediani, J. D., Ward, R. J., Milligan, G. (2017) A molecular basis for selective antagonist destabilization of dopamine D3 receptor quaternary organization. Scientific Reports, 7, (doi: 10.1038/s41598-017-02249-3)
Carmody, R., Keeshan, K. (2017) The Tribble with APL: a new road to therapy. Cancer Cell, 31, pp. 612-613. (doi: 10.1016/j.ccell.2017.04.011)
Houthuijzen, J. M., Oosterom, I., Hudson, B. D., Hirasawa, A., Daenen, L. G.M., McLean, C. M., Hansen, S. V.F., Jaarsveld, M. T.M., Peeper, D. S., Sadatmand, S. J., Roodhart, J. M.L., van de Lest, C. H.A., Ulven, T., Ishihara, K., Milligan, G., Voest, E. E. (2017) Fatty acid 16:4(n-3) stimulates a GPR120-induced signaling cascade in splenic macrophages to promote chemotherapy resistance. FASEB Journal, 31, pp. 2195-2209. (doi: 10.1096/fj.201601248R)
Papanatsiou, M., Amtmann, A., Blatt, M. R. (2017) Stomatal clustering in Begonia associates with the kinetics of leaf gaseous exchange and influences water use efficiency. Journal of Experimental Botany, 68, pp. 2309-2315. (doi: 10.1093/jxb/erx072)
Tanaka, M., Ishizuka, K., Nekooki-Machida, Y., Endo, R., Takashima, N., Sasaki, H., Komi, Y., Gathercole, A., Huston, E., Ishii, K., Hui, K. K.-W., Kurosawa, M., Kim, S.-H., Nukina, N., Takimoto, E., Houslay, M. D., Sawa, A. (2017) Aggregation of scaffolding protein DISC1 dysregulates phosphodiesterase 4 in Huntington’s disease. Journal of Clinical Investigation, 127, pp. 1438-1450. (doi: 10.1172/JCI85594)
Brandani, G. B., Vance, S. J., Schor, M., Cooper, A., Kennedy, M. W., Smith, B. O., MacPhee, C. E., Cheung, D. L. (2017) Adsorption of the natural protein surfactant Rsn-2 onto liquid interfaces. Physical Chemistry Chemical Physics, 19, pp. 8584-8594. (doi: 10.1039/C6CP07261E)
Parnell, E., McElroy, S. P., Wiejak, J., Baillie, G. L., Porter, A., Adams, D. R., Rehmann, H., Smith, B. O., Yarwood, S. J. (2017) Identification of a novel, small molecule partial agonist for the cyclic AMP sensor, EPAC1. Scientific Reports, 7, (doi: 10.1038/s41598-017-00455-7)
Ahmed, A., Wan, X., Mitxitorena, I., Lindsay, A. J., Pandolfi, P. P., McCaffrey, M. W., Keeshan, K., Chen, Y. H., Carmody, R. J. (2017) Regulation of NF-κB by PML and PML-RARα. Scientific Reports, 7, (doi: 10.1038/srep44539)
Chen, S.C., Brooks, R., Houskeeper, J., Bremner, S.K., Dunlop, J., Viollet, B., Logan, P.J., Salt, I.P., Ahmed, S.F., Yarwood, S.J. (2017) Corrigendum to "Metformin suppresses adipogenesis through both AMP-activated protein kinase (AMPK)-dependent and AMPK-independent mechanisms" [Mol. Cell. Endocrinol. 440 15 January 2017 57-68] Molecular and Cellular Endocrinology, 443, pp. 176. (doi: 10.1016/j.mce.2017.01.049)
Chen, Z.-H., Chen, G., Dai, F., Wang, Y., Hills, A., Ruan, Y.-L., Zhang, G., Franks, P. J., Nevo, E., Blatt, M. R. (2017) Molecular evolution of grass stomata. Trends in Plant Science, 22, pp. 124-139. (doi: 10.1016/j.tplants.2016.09.005)
Mackenzie, A. E., Milligan, G. (2017) The emerging pharmacology and function of GPR35 in the nervous system. Neuropharmacology, 113, pp. 661-671. (doi: 10.1016/j.neuropharm.2015.07.035)
Mancini, S. J., White, A. D., Bijland, S., Rutherford, C., Graham, D., Richter, E. A., Viollet, B., Touyz, R. M., Palmer, T. M., Salt, I. P. (2017) Activation of AMP-activated protein kinase rapidly suppresses multiple pro-inflammatory pathways in adipocytes including IL-1 receptorassociated kinase-4 phosphorylation. Molecular and Cellular Endocrinology, 440, pp. 44-56. (doi: 10.1016/j.mce.2016.11.010)
Chen, S. C., Brooks, R., Houskeeper, J., Bremner, S. K., Dunlop, J., Viollet, B., Logan, P. J., Salt, I. P., Ahmed, S. F., Yarwood, S. J. (2017) Metformin suppresses adipogenesis through both AMP-activated protein kinase (AMPK)-dependent and AMPK-independent mechanisms. Molecular and Cellular Endocrinology, 440, pp. 57-68. (doi: 10.1016/j.mce.2016.11.011)
Karnik, R., Waghmare, S., Zhang, B., Larson, E., Lefoulon, C., Gonzalez, W., Blatt, M. R. (2017) Commandeering channel voltage sensors for secretion, cell turgor, and volume control. Trends in Plant Science, 22, pp. 81-95. (doi: 10.1016/j.tplants.2016.10.006)
Zhang, B., Karnik, R., Waghmare, S., Donald, N. A., Blatt, M. R. (2017) VAMP721 conformations unmask an extended motif for K+ channel binding and gating control. Plant Physiology, 173, pp. 536-551. (doi: 10.1104/pp.16.01549)
Milligan, G., Shimpukade, B., Ulven, T., Hudson, B. D. (2017) Complex pharmacology of free fatty acid receptors. Chemical Reviews, 117, pp. 67-110. (doi: 10.1021/acs.chemrev.6b00056)
Hansen, A. H., Sergeev, E., Pandey, S. K., Hudson, B. D., Christiansen, E., Milligan, G., Ulven, T. (2017) Development and characterization of a fluorescent tracer for the free fatty acid receptor 2 (FFA2/GPR43) Journal of Medicinal Chemistry, 60, pp. 5638-5645. (doi: 10.1021/acs.jmedchem.7b00338)
(2017) Free Fatty Acid Receptors. 236, (doi: 10.1007/978-3-319-50693-7)
Milligan, G., Bolognini, D., Sergeev, E. (2017) Ligands at the free fatty acid receptors 2/3 (GPR43/GPR41) Springer International Publishing
Postgraduate research students
Refine By
-
{{student.surname}} {{student.forename}}
{{student.surname}} {{student.forename}}
({{student.subject}})
{{student.title}}
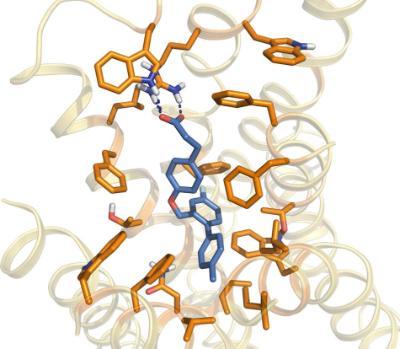 Molecular Pharmacology and Cellular Signalling are core themes with our School. Focus is on the structure, function and regulation of members of the family of G protein-coupled receptors (GPCRs) and how information deriving from detailed analysis of their mechanisms of action can be translated into the development of novel medicines and approaches to both understand and ameliorate disease. Underpinned for many years by the work of Professor Graeme Milligan, the Molecular Pharmacology Group is currently undergoing substantial and rapid expansion.
Molecular Pharmacology and Cellular Signalling are core themes with our School. Focus is on the structure, function and regulation of members of the family of G protein-coupled receptors (GPCRs) and how information deriving from detailed analysis of their mechanisms of action can be translated into the development of novel medicines and approaches to both understand and ameliorate disease. Underpinned for many years by the work of Professor Graeme Milligan, the Molecular Pharmacology Group is currently undergoing substantial and rapid expansion.
We use approaches ranging from classical pharmacological analyses to the development of biosensors based on biophysical methods and, linked to the generation of unique animal models to study disease development and the molecular mechanisms that underpin the basis of physiology, novel assay technologies allow better discrimination between ligand chemotypes to prioritise selection of candidate molecules for lead optimisation and development.
Close links have been established with both clinicians and basic science groups in other Schools within the College of Medical, Veterinary & Life Sciences with focus on cardiovascular and metabolic health, infection and immunity and dementia and neurodegeneration.


