Emulation & uncertainty quantification
We develop emulation methods and computationally tractable statistical models of complex processes such as mathematical models and computational digital twins.
Staff
Postgraduate research students
Refine By
-
{{student.surname}} {{student.forename}}
{{student.surname}} {{student.forename}}
({{student.subject}})
{{student.title}}
Emulation and Uncertainty Quantification - Example Research Projects
Information about postgraduate research opportunities and how to apply can be found on the Postgraduate Research Study page. Below is a selection of projects that could be undertaken with our group.
Structured ML for physical systems (PhD or MSc)
Supervisors: Lawrence Bull
Relevant research groups: Machine Learning and AI, Emulation and Uncertainty Quantification
When using Machine Learning (ML) for science and engineering, an alternative mindset is required to build sensible representations from data. Unlike other applications (e.g. large language models), the datasets are relatively small and curated - i.e. they are collected via experiments rather than scraped from the internet. The limited variance of training data typically renders learning by ‘brute force’ infeasible. Instead, we must encode domain-specific knowledge within ML algorithms to enforce structure and constrain the space of possible models.
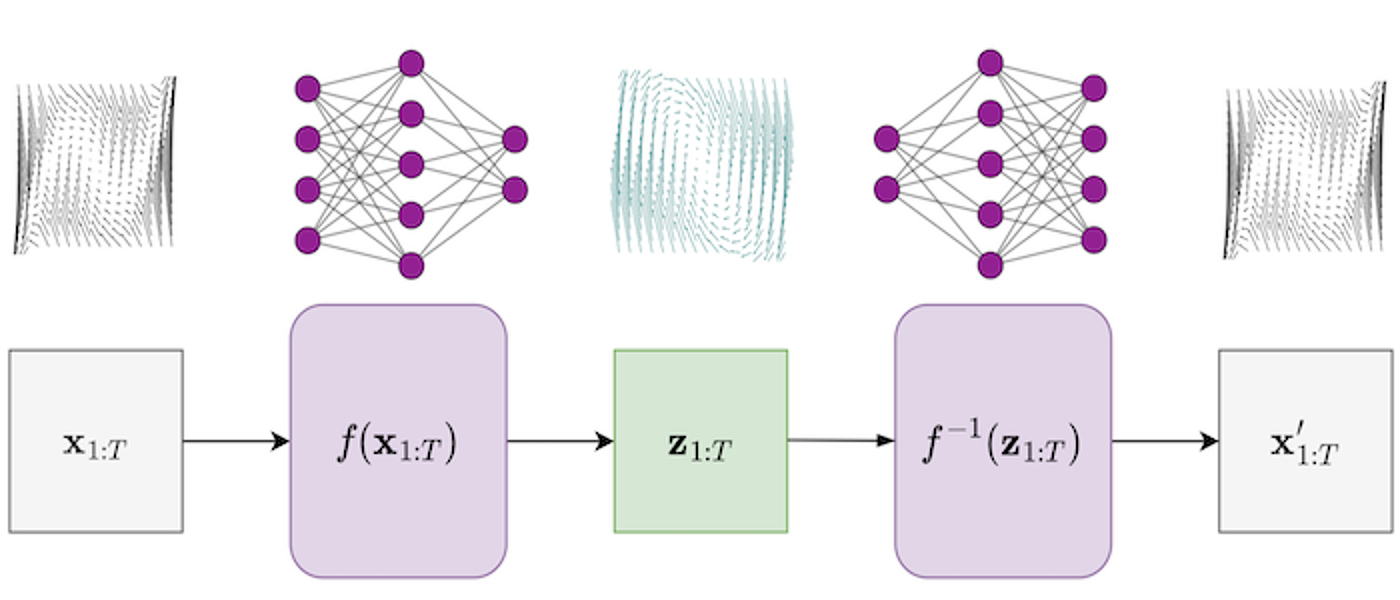
This project covers ML for physical systems (Karniadakis, 2021) and looks to integrate machine learning with applied mathematics - fusing scientific knowledge with insights from data. Methods will investigate various levels of constraints on ML predictions - including smoothness, invariances, etc. Relevant topics include:
- physics-informed machine learning (Girin, 2020)
- scientific machine learning (Pförtner, 2022)
- hybrid modelling
Seminars
Regular seminars relevant to the group are held as part of the Statistics seminar series. The seminars cover various aspects across the AI3 initiative and usually span multiple groups. You can find more information on the Statistics seminar series page, where you can also subscribe to the seminar series calendar.
Impressive advancements in mathematical modelling and digital twins (computer twins of complex systems) have highlighted a need for properly understanding parameter uncertainty and model calibration. In recent years, the Emulation and Uncertainty Quantification group has developed emulation methods, computationally tractable statistical models of complex processes, which help to infer parameters and uncertainty estimates in complex mathematical models.
The group collaborates heavily with the Continuum Mechanics groups within Applied Mathematics and leads parts of the SoftMech Centre for Healthcare in Mathematics. The group recently won the RSS Mardia prize to host workshops on environmental digital twins as part of the Analytics for Digital Earth project.