Professor Marshall Stark
My expertise relevant to Synthetic Biology is on enzymes that rearrange DNA sequences. Site-specific recombinases (SSRs) are enzymes that recognize target sequences (sites) in DNA molecules, bring two of the sites together, cut the DNA strands, and rejoin them in a new arrangement. SSRs have tremendous potential as tools for DNA sequence assembly, DNA micromanipulation, DNA-based counting systems and biosensors, and systematic genome modification of industrially important organisms. In my lab we are expanding the functionality of existing SSR systems and constructing novel systems by protein engineering.
I am the Principal award holder of a sLOLA grant from the BBSRC for the development of an SSR-based platform for Synthetic Biology. The aim is to use serine integrases to assemble and rearrange DNA sequences; this would allow us to engineer microbial cells so that they perform useful functions such as producing chemicals or degrading waste materials.
Current project:
A platform for rapid and precise DNA module rearrangements in Synthetic Biology.
Funded by BBSRC.
A team of researchers led by Professor Marshall Stark (IMCSB) has just been awarded a prestigious “sLOLA” (strategic, longer, larger) grant by the BBSRC. The 5-year grant (full value £4.0 Million) is for development of novel methods for splicing together and rearranging DNA sequences. The technology will involve enzymes called site-specific recombinases, which have been studied in Prof. Stark’s group for many years. These enzymes act as “molecular scissors and glue” for DNA, cutting the double helix at precisely defined positions and pasting new DNA segments into the gap. The major applications of the planned “SIDR” technology will be in the rapidly developing field of Synthetic Biology, which aims to engineer living organisms to produce useful substances such as industrial raw materials, biofuels, pharmaceuticals and food additives. The recombinases will be used to assemble strings of genes that encode enzymes needed to make the desired product, and will also allow subsequent modification of the genes to optimize product production levels. In another application of the technology, we hope to be able to make cells that can count, and keep a tally of the number they have counted up to in their DNA. This could allow us (for example) to record the number of times an individual cell has divided, or the number of times it has been exposed to a chemical substance.
This collaborative grant will fund research by Dr Sean Colloms, Prof Susan Rosser and Prof Stark at Glasgow, as well as groups in Aberdeen, York and Nottingham Universities led by Dr Oliver Ebenhoeh, Prof. Maggie Smith, and Dr. william Brown respectively. Two industrial partners, Ingenza Ltd. and Biotica Ltd., will also make important contributions to development of the project and commercialization of the research outputs. The funding is part of a £20M Government initiative aiming to make the UK a world leader in Synthetic Biology research and applications.
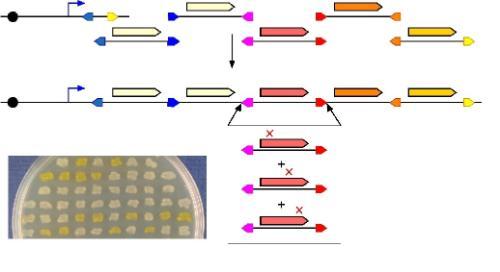
[Picture: Our SIDR technology can assemble individual gene cassettes (coloured long arrows) into arrays and then exchange individual cassettes with variant versions (mutations marked with red crosses), by recombination between matched pairs of sites (coloured short arrows). The thin blue bent arrow indicates a promoter. The inset shows E. coli strains which have been engineered in this way, to produce different levels of the yellow pigment Zeaxanthin.]