Prof Barbara Mable
Prof Barbara Mable
 Professor of Evolutionary Genetics
Professor of Evolutionary Genetics
Institute of Biodiversity, Animal Health & Comparative Medicine
College of Medicine, Veterinary & Life Sciences
Graham Kerr Building
University of Glasgow
Glasgow, G12 8QQ
Tel.: +44 (0)141 330 3532
Email: Barbara.Mable@glasgow.ac.uk
Academic History
Academic History
- 2014-present Professor of Evolutionary Genetics, Institute of Biodiversity, Animal Health and Comparative Medicine, University of Glasgow
- 2011-2014 Reader, Institute of Biodiversity, Animal Health and Comparative Medicine, University of Glasgow
- 2007-2011 Senior Research Fellow, Division of Environmental and Evolutionary Biology, University of Glasgow
- 2005-2006 NERC Advanced Research Fellow, Division of Environmental and Evolutionary Biology, University of Glasgow
- 2004-2005 Lecturer, Division of Environmental and Evolutionary Biology, University of Glasgow
- 2000-2005 Assistant Professor, Department of Botany, University of Guelph
- 1998-2000 Postdoctoral Fellow, ICAPB, University of Edinburgh (BBSRC grant to Deborah Charlesworth)
- 1996-1998 Postdoctoral Fellow, Dept. of Zoology, University of British Columbia (Killam Memorial Fellowship to BKM and NSERC award to Sarah P. Otto)
- 1992-1996 Ph.D. (Zoology) University of Texas at Austin
- 1989-1991 Research Associate, Agriculture Canada Research Station Project: Genetics of resistance to acaricides in European Red Mites on apples.
- 1987-1989 M.Sc. (Zoology) University of Guelph
Fellowships
Fellowships
- Marie Curie Incoming International Fellowship, 2004-2006 (Declined)
- NERC Advanced Research Fellowship, University of Glasgow, 2004-2009
- NSERC University Faculty Award, University of Guelph, 2000-2005
- Izaak Walton Killam Memorial Fellowship, University of British Columbia, 1996-1998
Current Teaching Responsibilities
- Cluster Leader, Animal and Plant Sciences Postgraduate Taught Master’s (PGT) Programmes (since 2014)
- PGT Co-ordinator for programmes offered through the Institute of Biodiversity, Animal Health and Comparative Medicine (since 2016)
- Course Co-ordinator, Master’s Level: Key Research Skills; Conservation Genetics (since 2011)
- Course Co-ordinator, Undergraduate: L3 Applied Evolution (since 2018)
- Other undergraduate teaching: L2 Animal Biology, Evolution and Ecology (since 2018); L3 Tutorials (since 2004); L4 Evolution (since 2004)
Research Overview
Research in my laboratory (the evolutionary genetics group) is directed towards understanding how changes at the molecular level affect cellular and whole organism processes in a wide range of organisms (Figure 1). Most of my early work focused on the genetic and ecological consequences of a particularly extreme form of genetic change- whole genome duplication or polyploidy- but I have also been interested in the consequences of gene duplication at the level of gene families that control recognition processes involved in adaptation, such as plant self-incompatibility systems (SI), plant resistance genes, and the vertebrate immune genes (e.g. the Major Histocompatibility Complex, MHC). I am particularly interested in how such genomic changes affect interactions between organisms, such as mate choice and pathogen responses. Some of my primary areas of interest have been: 1) the evolutionary dynamics of gene families involved in recognition systems; 2) the causes and consequences of changes in mating systems for genetic diversity and adaptation; and 3) the consequences of mating system variation and polyploidy for mate recognition and pathogen response.
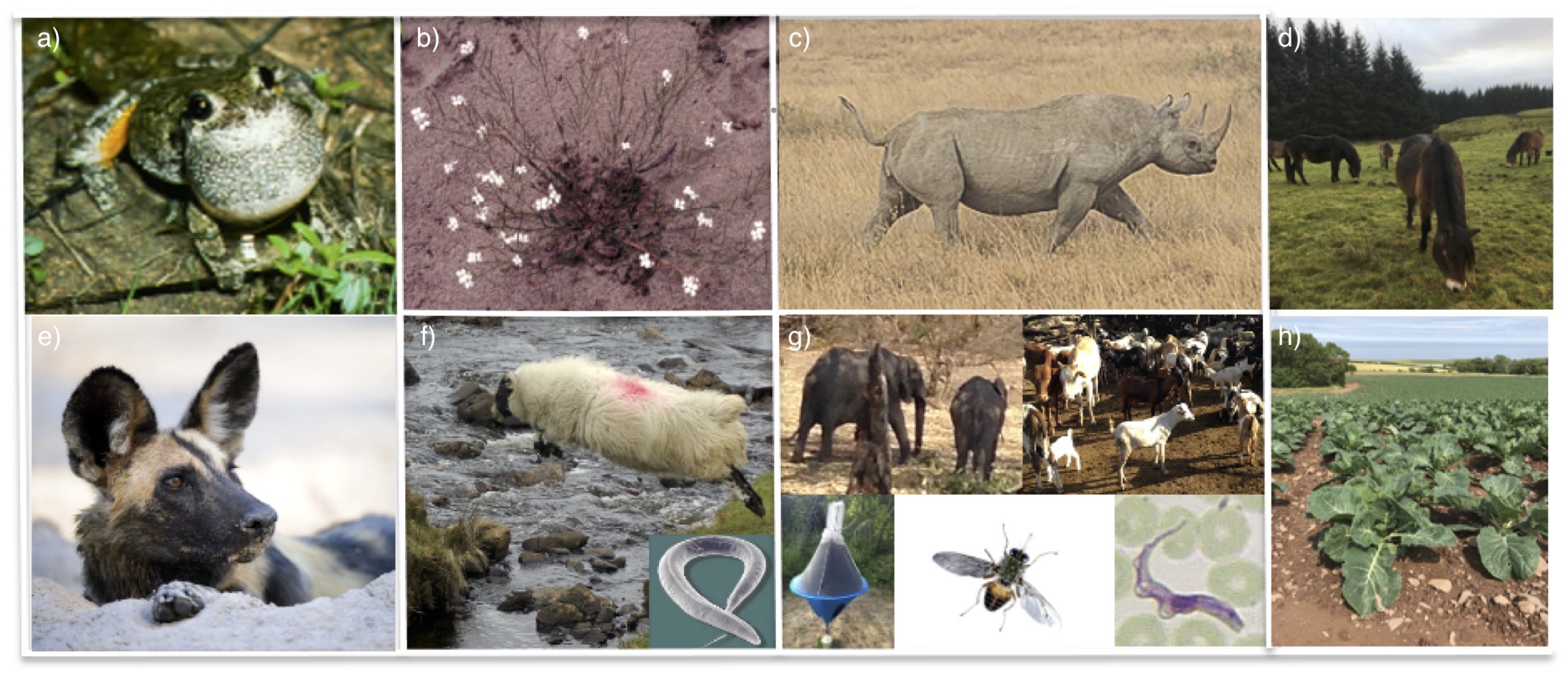

Figure 1. Some research questions and study organisms: evolution of mating systems and polyploidy in a) grey treefrogs (Hyla versicolor) and b) rock cress (Arabidopsis lyrata); conservation genetics of c) black rhinoceros, d) Exmoor ponies and e) African wild dogs; f) evolution of resistance to anthelmintics in nematodes infecting livestock; g) host-pathogen-vector dynamics (e.g. multiple species of tsetse flies feed on a wide range of wild and domestic hosts and can transmit multiple species of trypanosomes); and h) the influence of soil microbial communities on control of bacterial pathogens of crops.
Although we work on a wide range of organisms (bacteria, fungi, oomycetes, plants, nematodes, trematodes, kinetoplastids, molluscs, insects, amphibians, fish, birds and mammals), a central theme is considering the relative importance of genetic variation in general: how critical is it for adaptation and can genetics be used effectively to inform management interventions for conservation and disease mitigation? This has also led to new funding opportunities that have enabled diversification of our evolutionary genetics research team (Figure 2).
Figure 2. Lockdown teamwork: remote meeting of the evolutionary genetics group.
Ongoing projects led by recent or current PhD students include: 1) assessing diversity of the rhizosphere of plants in the genus Brassica in relation to cultivars and population genetic structure (Elizabeth Mittell, PhD awarded in 2019, funded by a Lord Kelvin and Adam Smith Scholarship); 2) developing molecular markers to inform translocation strategies for fenced populations of Southern White Rhinoceros in Botswana (Tarid Purisotayo, PhD awarded in 2020, funded by the government of Thailand); 3) investigating the genetic and ecological drivers of resistance in parasitic nematodes of sheep (Sam Brown, PhD candidate, funded by NERC); 4) conservation genetics and rewilding of Exmoor ponies (Debbie Davy, PhD candidate, funded by the Exmoor Pony Society); 5) genomic selection for immune function in cattle (Kamonchanok [June] Chooyoung, PhD candidate, funded by the government of Thailand); 6) assessing levels of genetic variation in protected populations of Northern Black Rhinoceros across Tanzania (Ronald Vincent Mellya, PhD candidate, funded by Zurich Zoo); and 7) how climate change is affecting economically important fish in Nigeria (Mark Kauna Sanda, PhD candidate, funded by a Commonwealth scholarship).
In collaboration with Umer Zeeshan Ijaz, Tom Morrison, Grant Hopcraft at Glasgow, and Linus Munishi at the Nelson Mandela African Institute of Science and Technology, a recent project led by two former master’s students, Koggani Koggani (funded by a Karimjee Jivanjee Scholarship and the Rufford Foundation) and Rachel Gray, is to use deep sequencing of chloroplast DNA amplicons to compare the diets of endangered ungulates and encroaching cattle populations.
Two funded collaborative research projects are led by postdoctoral fellows (Anubhab Khan and Ciara Keating) and critically supported by research technicians (Elizabeth Kilbride and Ryan Carter): 1) “What can you learn from a fly? Biting insects as noninvasive samplers” (funded by the Leverhulme Trust), in collaboration with Harriet Auty (IBAHCM) and a former Phd student, Manun Channumsin; and 2) “A Decision Support tool for Potato Blackleg Disease (DeS-BL)” (funded by BBSRC, NERC, DEFRA and the Scottish government), a consortium grant in the Bacterial Plant Diseases Programme, led by the James Hutton Institute (Ian Toth PI), in collaboration with Umer Zeeshan Ijaz (School of Engineering) and Joel Milner (MCSB) at Glasgow; you can hear Ciara talk about the project here.
Current Research
Most of the research in my laboratory is currently focused on understanding the adaptive dynamics of plants in variable environments. In particular, I am interested in the effects of mating systems (i.e. inbreeding vs outcrossing) and ploidy level (i.e. diploid vs tetraploid) on the ability of plants to tolerate or adapt to changes in the abiotic and the biotic (e.g. changes in pathogen pressures or proximity to competitors) environments.
We have been using A. lyrata as a model to investigate how mating system and polyploidy affect pathogen response systems. We have been focusing on particular oomycete pathogens (causing white blister rust and downy mildew) that are known to be problematic for cultivated species of Brassicaceae and for which candidate genes for resistance have been previously characterised in Arabidopsis thaliana. We have been investigating variation in resistance to pathogens in plants sampled from across their European range (where the plants are known to vary in ploidy level) and in the Great Lakes region of Eastern North America (where the plants are known to vary in mating system) and investigating the prevalence of the pathogens in native environments and variation in candidate resistance genes. We have also been using RAD sequencing to enable comparison of genome-wide patterns of variation in neutral genes with those that might be involved in adaptation. This research is currently funded by a grant from NERC, in collaboration with Eric Holub, from the University of Warwick. The project has involved three postdoctoral researchers (Volkan Cevik and Joana Vicente, Warwick; James Buckley, Glasgow), and three research technicians (Aileen Adam, Elizabeth Kilbride, Ryan Carter).
We are also using the University field station on Loch Lomond (SCENE) to pilot use of outdoor experimental nurseries to use a common garden garden approach to investigate the ability of plants with different ploidy and mating systems to adapt to novel and variable environmental conditions (Figure 2).
 |
 |
Figure 2. Experimental nursery at SCENE showing natural variation in climatic conditions.

