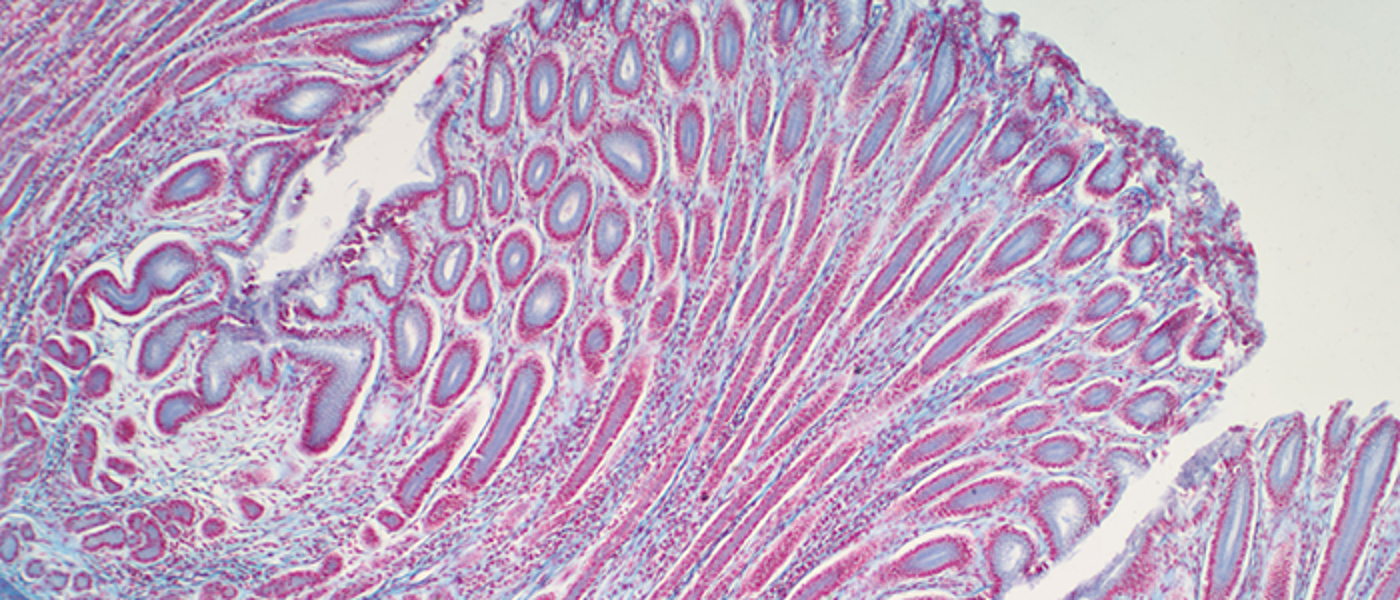
Colorectal/Lower GI/intestinal/gut microbiome
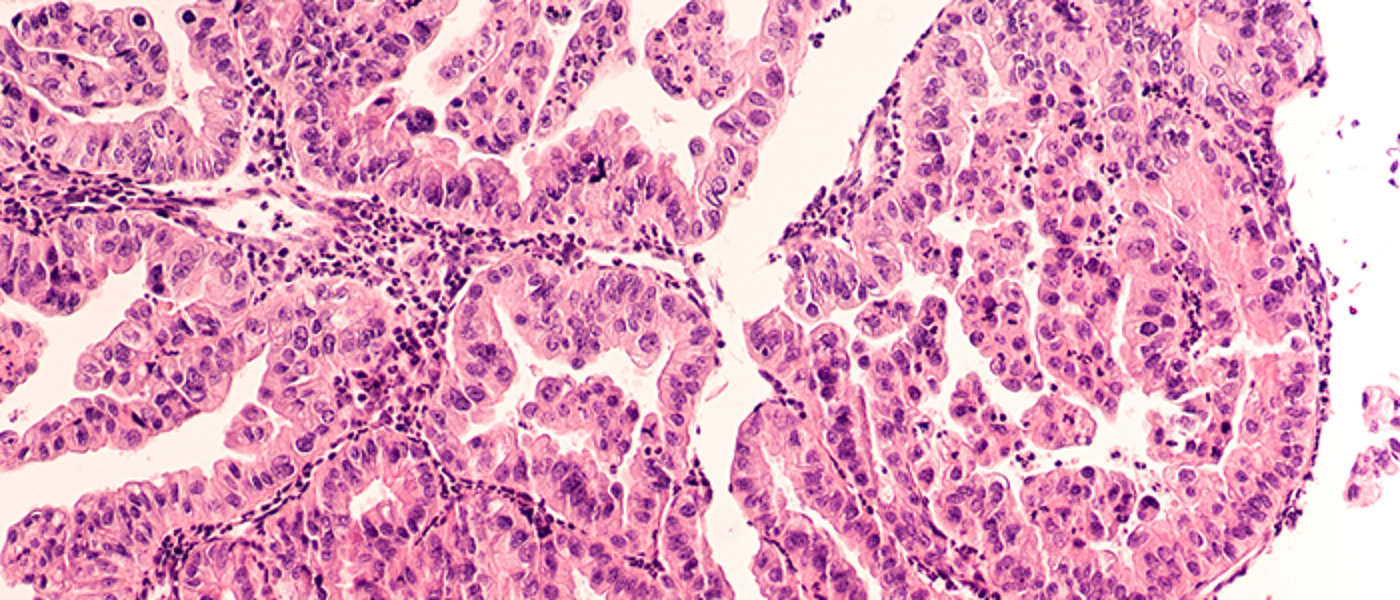
Gynaecological (uterine/ovarian)

Upper GI (cholangiocarcinoma/biliary cancers/hepatobiliary)
Urological (Prostate/Bladder/Kidney)
Research focused on the most common form of cancer for men in the UK.












