A New Auto-Inductive Chemical System for Indirect Protein Sensing
Supervisor: Dr Liam Wilson
School: Chemistry
Description:
Sensitive and specific protein sensing is essential in clinical diagnostics and environmental/food monitoring. Gold-standard assays typically utilise antibodies as recognition motifs for target proteins (e.g. ELISA and lateral-flow immunoassays). However, reliance on antibody reagents can introduce practical challenges: high reagent cost, batch-to-batch variability, cold-chain storage requirements, and limited reagent stability/shelf life. As such, there is demand for operationally simple protein-sensing methodologies that circumvent the need for antibodies while retaining sensitivity and specificity.
We aim to develop an assay that indirectly senses target proteins using small-molecule ‘turn-on’ fluorescent probes, via a two-step process (Figure 1A). The core concept is to use a strong, selective non-covalent ligand for the target protein, which bears a sulfur(VI) fluoride exchange (SuFEx) motif. This motif will undergo proximity-driven reaction with nearby nucleophilic amino-acid side-chains, releasing fluoride. We propose that the released fluoride can be amplified and detected using probes such as DPF-1 (Figure 1B). Activation of DPF-1 by a fluoride input triggers fluorescence and releases two additional equivalents of fluoride, creating a positive-feedback (‘auto-inductive’) loop. This should enable detection of very low initial fluoride concentrations (pM), translating into ultrasensitive indirect protein sensing.
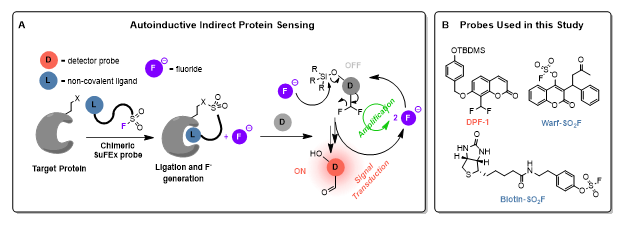
This project will establish a proof-of-concept for this sensing platform. We will target two proteins: streptavidin (Strp) and human serum albumin (HSA), using SuFEx analogues of well-characterised ligands (biotin and warfarin, respectively): Biotin-SO2F and Warf-SO2F (Figure 1B). Samples of DPF-1, Biotin-SO2F and Warf-SO2F have been prepared in-house.
The student’s objectives are to:
- Optimise fluoride amplification and detection. The student will incubate DPF-1 with varying concentrations of fluoride standards, systematically varying solvent, base, and temperature. Reactions will be monitored by plate-based fluorimetry to optimise the fluoride limit of detection (LOD).
- Investigate target binding and ligation of Biotin-SO2F and Warf-SO2F, over time, via intact-protein LC-MS, and examine background hydrolysis of the probes via F NMR. These data will be used to optimise probe-protein incubation times and minimize non-productive fluoride release.
- Demonstrate indirect protein detection. Using the optimised conditions, the student will perform protein-sensing experiments across a range of Strp and HSA concentrations, using Biotin-SO2F and Warf-SO2F, followed by DPF-1, via plate-based fluorimetry. Controls (e.g. ligand without SuFEx, protein-free blanks, and fluoride standards) will validate the sensing pathway and allow protein LODs to be established.

