CVR Genomics
High-throughput sequencing is a vital tool for scientific research and to guide public health actions. CVR Genomics is a team of researchers exclusively dedicated to the development and implementation of high-throughput sequencing solutions applied to viruses. Working in close alignment with the CVR’s Bioinformatics team, we support CVR researchers, external collaborators and provide resources to the virology community.
CVR Genomics’ cutting-edge equipment and expertise supports the study of viruses of human and veterinary relevance. Our know-how has been acquired through experimental troubleshooting, with over 240 viruses sequenced, such as blood and respiratory viruses, arboviruses and viral haemorrhagic viruses, across different host species.
The facility runs the following sequencing platforms:
- Illumina
- Oxford Nanopore Technologies
And the following sequencing pipelines:
- Viral genomics
- Metagenomics
- Amplicon
- Probe-based capture
- Transcriptomics
- mRNA
- Total RNA
- Small RNA
- Single-cell omics
Sharing know-how is important to propel scientific output, hence the facility provides customised training and advice to help deliver specific projects.
Due to its collective expertise and associated infrastructure, CVR Genomics is well positioned to respond promptly to viral threats and has been involved in the development and implementation of protocols to support surveillance, viral discovery and pandemic response.
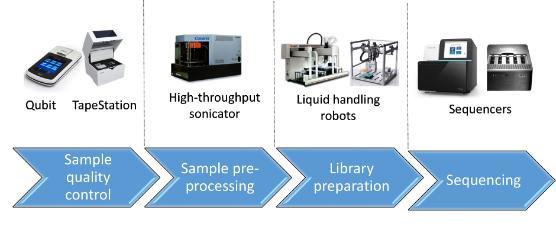
CVR Genomics houses an array of specialised equipment, covering different stages of the sequencing process.
Through internal collaborations with CVR researchers and external collaborations with public health agencies, the NHS, veterinary labs, and other research institutes, we aim to continue implementing bespoke/innovative sequencing approaches for the study of viruses and virus-host interactions.
More information about our equipment
Illumina MiSeq
 The MiSeq is a fully-integrated benchtop Illumina sequencer that features a short run time and highly accurate sequencing chemistry.
The MiSeq is a fully-integrated benchtop Illumina sequencer that features a short run time and highly accurate sequencing chemistry.
The MiSeq is ideal for sequencing small genomes (such as viruses and bacteria), variant discovery and mutation detection, PCR amplicon sequencing, ChIP-Seq, gene expression analysis, small RNA discovery and metagenomics. Up to 96 samples can be multiplexed on one run using dual index tags. The maximum output is currently 9 Gb (9,000,000,000 bp) and up to 30 million reads.
Ion Proton Sequencer
 The Ion Proton DNA sequencer is a semiconductor-based benchtop instrument that has sufficient power for human-scale genome, exome and whole transcriptome sequencing.
The Ion Proton DNA sequencer is a semiconductor-based benchtop instrument that has sufficient power for human-scale genome, exome and whole transcriptome sequencing.
Agilent Tapestation
 The Agilent TapeStation is replacing the Bioanalyser for automated sample QC. Electrophoresis is carried out on polymer tracks called ScreenTapes which are available for DNA, RNA and proteins.
The Agilent TapeStation is replacing the Bioanalyser for automated sample QC. Electrophoresis is carried out on polymer tracks called ScreenTapes which are available for DNA, RNA and proteins.
Each ScreenTape has 16 analysis tracks that can either all be used at the same time, or alternatively a minimum of 2 lanes can be run independently. Complete analysis of 16 samples takes approximately 20 minutes.
Covaris S220 Sonicator
 The Covaris sonicator uses adaptive focused acoustics to shear DNA into specific fragment lengths for generating next-generation sequencing libraries.
The Covaris sonicator uses adaptive focused acoustics to shear DNA into specific fragment lengths for generating next-generation sequencing libraries.
The Sonicator can be used for DNA, RNA, and chromatin shearing, tissue homogenization, cell lysis, compound dissolution, and particle micronization.
Bursts of ultrasonic energy are focused into the sample vessel which is immersed in a cooled water bath to prevent heat damage. Different sample tubes are available to allow volumes from 25ul to 10ml to be sheared to specific fragment lengths.
3 Mosquito® HV
 This is a nanodispenser that uses positive displacement, enabling the reduction in reagent volumes for library preparation. It offers highly accurate and precise multichannel pipetting from 500 nL to 5 µL, leading to significant savings on precious reagents.
This is a nanodispenser that uses positive displacement, enabling the reduction in reagent volumes for library preparation. It offers highly accurate and precise multichannel pipetting from 500 nL to 5 µL, leading to significant savings on precious reagents.
GridION sequencer
 This is a flexible sequencer that processes DNA and RNA molecules using nanopores, with sequence lengths corresponding to the length of the molecules being sequenced. It has a throughput of as much as 150 Gb of data thanks to the five Flongle or MinION flow cells it can run simultaneously, which is streamed in real time for immediate analysis. It has a powerful data processing and analysis capacity, minimising IT requirements.
This is a flexible sequencer that processes DNA and RNA molecules using nanopores, with sequence lengths corresponding to the length of the molecules being sequenced. It has a throughput of as much as 150 Gb of data thanks to the five Flongle or MinION flow cells it can run simultaneously, which is streamed in real time for immediate analysis. It has a powerful data processing and analysis capacity, minimising IT requirements.

