Research
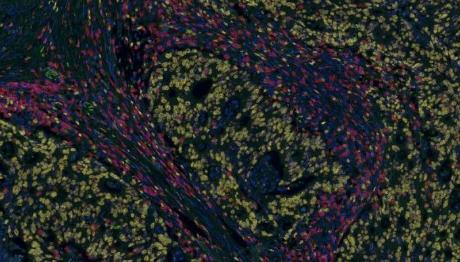
The main aim of the Translational Cancer Pathology Group is to better understand the determinants of short- and long-term outcomes in patients with colorectal cancer.
We are interested in the relationship between local and systemic inflammatory responses, tumour microenvironment and cell signalling with a view to taking such knowledge into the routine clinical management of colorectal cancer.
Our overarching aim is to develop novel treatment strategies to establish a personalised medicine approach for colorectal cancer.
Colorectal cancer
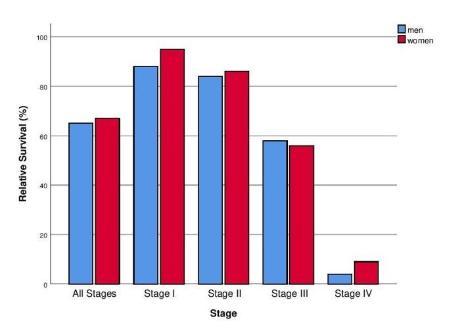 Colorectal cancer (CRC) poses a significant burden on worldwide healthcare, as it currently ranks third for cancer incidence and second for cancer related mortality. Despite use of multidisciplinary management and improved treatment, nearly half of all CRC patients present with metastasis either at the time of diagnosis or as recurrent disease. The metastatic process is one of the main contributing factors for the majority of CRC mortalities.
Colorectal cancer (CRC) poses a significant burden on worldwide healthcare, as it currently ranks third for cancer incidence and second for cancer related mortality. Despite use of multidisciplinary management and improved treatment, nearly half of all CRC patients present with metastasis either at the time of diagnosis or as recurrent disease. The metastatic process is one of the main contributing factors for the majority of CRC mortalities.
Currently the TNM staging system is the primary tool used for the management of CRC. However, this is less than ideal as it cannot account for the wide variation in prognosis that is observed at each disease stage, even when combined with markers such as MMR status and KRAS status. One of the most challenging steps to improve CRC treatment and survival is therefore identification of prognostic biomarkers to predict which patients are most at risk of recurrence and metastasis, alongside predictive biomarkers to predict therapeutic strategy at an individual patient level.
Professor Joanne Edwards and her team have therefore developed large tissue cohorts with linked clinical information to further investigate the molecular mechanisms underlying the development and progression of CRC. Through this work, Professor Edwards hopes to identify novel prognostic biomarkers and therapeutic targets for future cancer treatments.
Patient cohorts
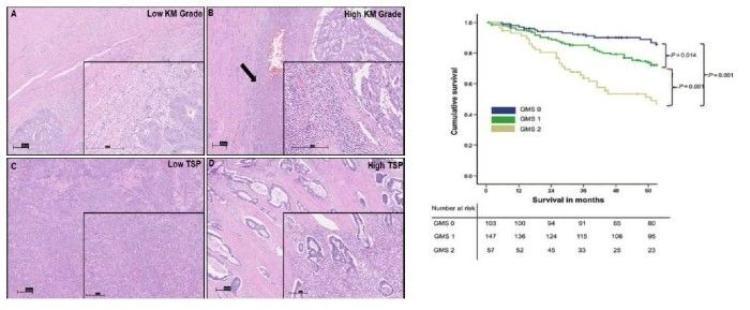 In order to study the molecular mechanisms underlying the development and progression of colorectal cancer, several different tissue cohorts have been developed with linked clinical data.
In order to study the molecular mechanisms underlying the development and progression of colorectal cancer, several different tissue cohorts have been developed with linked clinical data.
- INCISE cohort: 3000 retrospective precancerous colonic polyps.
- DM cohort: 142 retrospective stage I-III colorectal cancer.
- KP cohort: 1030 retrospective stage I-IV colorectal cancers.
- GRI cohort: 823 retrospective stage I-III colorectal cancer.
- Synchronous cohort: 44 stage IV colorectal cancer patients who underwent synchronous resection of primary colorectal tumours and matched liver metastases.
Technologies Employed

Tissue microarrays
Working in close collaboration with the Glasgow Tissue Research Facility (GTRF) and the NHS Greater Glasgow and Clyde Biorepository (NHSGG&C), we construct custom tissue microarrays (TMAs) to ensure maximal use of valuable clinical tissue.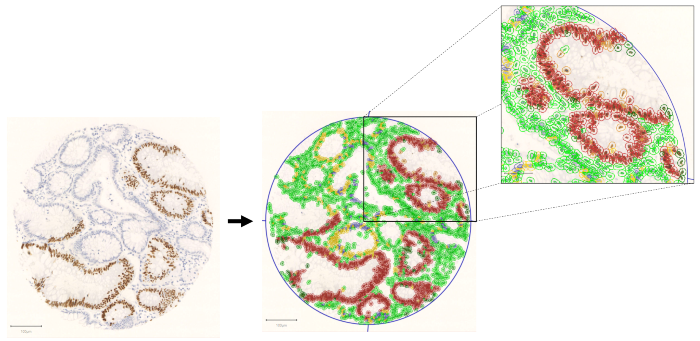
Digital pathology
Machine and deep learning methods are used to develop bespoke digital pathology algorithms for analysing tissue samples. Digital pathology tools are also used to validate manual scoring.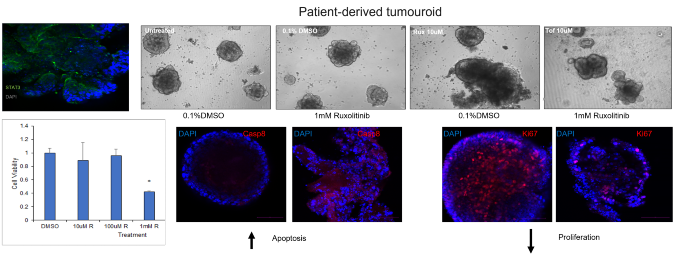
Patient-derived organoids
Patient-derived organoids are cultured in the laboratory and used to explore and validate novel therapeutic targets.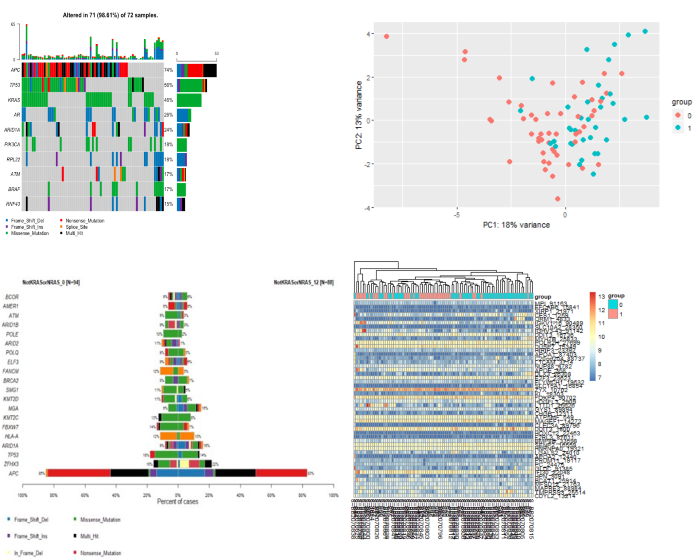
-omics technologies
A range of -omics technologies are used by our team to determine the somatic mutational landscape and transcriptomic profile in our tissue samples.